

Gibson assembly is a molecular cloning method that allows for the joining of multiple DNA fragments in a single, isothermal reaction. It is named after its creator, Daniel G. Gibson, who is the chief technology officer and co-founder of the synthetic biology company, Telesis Bio. The technology is more efficient than manual plasmid genetic recombination methods, but remains expensive as it is still under patent.[1][2]
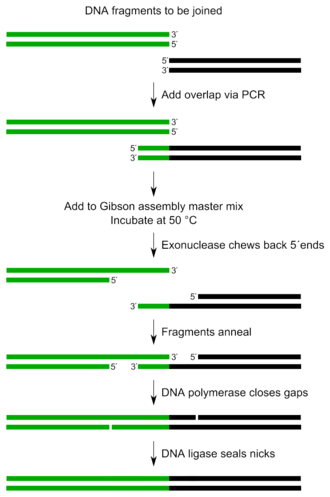
The entire Gibson assembly reaction requires few components with minor manipulations.[3]
The method can simultaneously combine up to 15 DNA fragments based on sequence identity. It requires that the DNA fragments contain ~20-40 base pair overlap with adjacent DNA fragments. These DNA fragments are mixed with a cocktail of three enzymes, along with other buffer components.
The three required enzyme activities are: exonuclease, DNA polymerase, and DNA ligase.
The resulting product is different DNA fragments joined into one. Either linear or closed circular molecules can be assembled.
There are two approaches to Gibson assembly. A one-step method and a two-step method. Both methods can be performed in a single reaction vessel. The Gibson assembly 1-step method allows for the assembly of up to 5 different fragments using a single step isothermal process. In this method, fragments and a master mix of enzymes are combined and the entire mixture is incubated at 50 °C for up to one hour. For the creation of more complex constructs with up to 15 fragments, or for constructs incorporating fragments from 100 bp to 10 kb, the Gibson assembly two-step approach is used. The two-step reaction requires two separate additions of master mix. One of the reactions is for the exonuclease and annealing step while the other is for DNA polymerase and ligation steps. For the two-step approach, different incubation temperatures are used to carry out the assembly process.
The Gibson DNA assembly method has many advantages compared to conventional restriction enzyme/ligation cloning of recombinant DNA. For example,
Gibson says he no longer ev en bothers with standard restriction enzyme-based cloning in his lab.