

| |
| Names | |
|---|---|
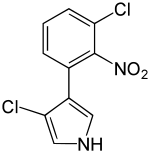
| Preferred IUPAC name
3-Chloro-4-(3-chloro-2-nitrophenyl)-1H-pyrrole | |
| Identifiers | |
3D model (JSmol) |
|
| ChEBI | |
| ChemSpider |
|
| ECHA InfoCard | 100.012.557 |
| EC Number |
|
| KEGG |
|
| MeSH | D011764 |
PubChem CID |
|
| UNII | |
CompTox Dashboard (EPA) |
|
| |
| |
| Properties | |
| C10H6Cl2N2O2 | |
| Molar mass | 257.07284 |
| Pharmacology | |
| D01AA07 (WHO) | |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
Pyrrolnitrin (PRN[1]) is a phenylpyrrole chemical used an antifungal antibiotic.[2] Pseudomonas pyrrocinia and other Pseudomonas species produce pyrrolnitrin from tryptophan as secondary metabolite.[3][4] It is believed that the antifungal properties come from inhibition of electron transport system.[5]
The synthetic fungicides fenpiclonil and fludioxonil are chemically related to pyrrolnitrin.[6][7]
InPseudomonas fluorescens, biosynthesis of pyrrolnitrin requires four genes, named prnABCD, arranged into a single operon. The products of these genes are similar in size and catalyze four subsequent reactions:[1][5]

Except for prnA, these enzymes are unable to act on D-tryptophan.[1][5]
Neither of the chlorinating enzymes, prnA nor prnC, show homology to known haloperoxidases nor to one another.[1]
An alternative pathway was also suggested, where L-tryptophan is first turned into aminophenylpyrrole (APP) and then by subsequent steps to aminopyrrolnitrin and pyrrolnitrin. While these steps have not been described in more detail, prnB is able to produce APP, presumably from tryptophan as starting material.[1] APP seems to be an unwanted side product. The gene coding for prnB also starts with the unusual GTG start codon, further lowering the amount of prnB expressed and thus lowering the amount of present APP.