

|
link backbone
|
|
||
| (29 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
{{dashboard.wikiedu.org sandbox}} |
{{dashboard.wikiedu.org sandbox}} |
||
[[File:Bruker Amazon Speed ETD.jpg|thumb|An ion trap mass spectrometer with electron transfer dissociation capability|320x320px]] |
|||
''Editing electron-transfer dissociation page for LSU Mass Spec class (Chem 4558) Spring 201'' |
|||
| ⚫ | [[File:Peptide fragmentation.gif|thumb|320x320px|Peptide fragmentation notation]] |
||
| ⚫ | '''Electron-transfer dissociation''' ('''ETD''') is a method of [[Fragmentation (mass spectrometry)|fragmenting]] multiply-charged gaseous [[macromolecule]]s in a [[mass spectrometer]] between the stages of [[tandem mass spectrometry]] (MS/MS).<ref>{{Cite book|title=Fundamentals of Contemporary Mass Spectrometry|last=Dass|first=Chhabil|publisher=John Wiley & Sons.|year=2007|isbn=978-0-470-11848-1|location=Hoboken, New Jersey|pages=128}}</ref> Similar to [[electron-capture dissociation]], ETD induces fragmentation of large, multiply-charged [[cations]] by transferring [[electron]]s to them.<ref name=":2">{{Cite journal|last=Hart-Smith|first=Gene|date=2014-01-15|title=A review of electron-capture and electron-transfer dissociation tandem mass spectrometry in polymer chemistry|url=http://www.sciencedirect.com/science/article/pii/S000326701301235X|journal=Analytica Chimica Acta|series=Polymer Mass Spectrometry|volume=808|pages=44–55|doi=10.1016/j.aca.2013.09.033}}</ref> ETD is used extensively with polymers and biological molecules such as [[protein]]s and [[peptide]]s for [[sequence analysis]].<ref name=":1">{{Cite journal|last=Brodbelt|first=Jennifer S.|date=2015-12-11|title=Ion Activation Methods for Peptides and Proteins|url=http://pubs.acs.org/doi/abs/10.1021/acs.analchem.5b04563|journal=Analytical Chemistry|language=EN|volume=88|issue=1|pages=30–51|doi=10.1021/acs.analchem.5b04563}}</ref> Transferring an electron causes [[Structure#Biological|peptide backbone]] cleavage into [[Peptide sequence tag|c- and z-ions]] while leaving [[Lability|labile]] [[Post-translational modification|post translational modifications]] (PTM) intact.<ref>{{Cite journal|last=Coon|first=Joshua J.,|last2=Syka|first2=John E.P.|last3=Shabanowitz|first3=Jeffrey|last4=Hunt|first4=Donald F.|date=April 2005|title=Tandem Mass Spectrometry for Peptide and Protein Sequence Analysis|url=http://www.biotechniques.com/BiotechniquesJournal/2005/April/Tandem-Mass-Spectrometry-for-Peptide-and-Protein-Sequence-Analysis/biotechniques-45438.html|journal=BioTechniques|doi=|pmid=15884666|access-date=April 15, 2016}}</ref> The technique only works well for higher charge state ions (z>2).<ref name=":2" /> However, relative to [[collision-induced dissociation]] (CID), ETD is advantageous for the fragmentation of longer peptides or even entire proteins.<ref name=":3">{{Cite journal|last=Qi|first=Yulin|last2=Volmer|first2=Dietrich A.|date=2015-10-01|title=Electron-based fragmentation methods in mass spectrometry: An overview|url=http://onlinelibrary.wiley.com/doi/10.1002/mas.21482/abstract|journal=Mass Spectrometry Reviews|language=en|pages=n/a–n/a|doi=10.1002/mas.21482|issn=1098-2787}}</ref> This makes the technique important for [[top-down proteomics]].The method was developed by [[Donald F. Hunt|Hunt]] and coworkers at the [[University of Virginia]].<ref>{{US patent reference | number = 7534622 | y = 2009 | m = 05 | d = 19 | inventor = Donald F. Hunt, Joshua J. Coon, John E.P. Syka, Jarrod A. Marto | title = Electron transfer dissociation for biopolymer sequence mass spectrometric analysis}}</ref> |
||
[[File:Bruker Amazon Speed ETD.jpg|thumb|300x300px|Bruker Amazon Speed ETD at Louisiana State University]] |
|||
| ⚫ |
'''Electron-transfer dissociation''' (ETD) is a method of [[Fragmentation (mass spectrometry)|fragmenting]] |
||
== History == |
== History == |
||
Electron-capture dissociation (ECD) was developed in 1998 to fragment large proteins for mass spectrometric analysis.<ref>{{Cite journal|last=Zubarev|first=Roman A.|last2=Kelleher|first2=Neil L.|last3=McLafferty|first3=Fred W.|date=1998-04-01|title=Electron Capture Dissociation of Multiply Charged Protein Cations. A Nonergodic Process|url=http://dx.doi.org/10.1021/ja973478k|journal=Journal of the American Chemical Society|volume=120|issue=13|pages=3265–3266|doi=10.1021/ja973478k|issn=0002-7863}}</ref> Because ECD requires a large amount of near-thermal electrons (<0.2eV), originally it was used exclusively with [[Fourier transform ion cyclotron resonance|Fourier transform ion cyclotron resonance mass spectrometry]] (FTICR), the most expensive form of MS instrumentation.<ref>{{Cite journal|last=McLafferty|first=Fred W.|last2=Horn|first2=David M.|last3=Breuker|first3=Kathrin|last4=Ge|first4=Ying|last5=Lewis|first5=Mark A.|last6=Cerda|first6=Blas|last7=Zubarev|first7=Roman A.|last8=Carpenter|first8=Barry K.|date=2001-03-01|title=Electron capture dissociation of gaseous multiply charged ions by Fourier-transform ion cyclotron resonance|url=http://link.springer.com/article/10.1016/S1044-0305%2800%2900223-3|journal=Journal of the American Society for Mass Spectrometry|language=en|volume=12|issue=3|pages=245–249|doi=10.1016/S1044-0305(00)00223-3|issn=1044-0305}}</ref> Less costly options such as [[Quadrupole time-of-flight mass spectrometer|quadrupole time-of-flight]] (Q-TOF), [[quadrupole ion trap]] (QIT) and [[linear quadrupole ion trap]] (QLT) instruments used the more energy-intensive [[collision-induced dissociation]] method (CID), resulting in random fragmentation of peptides and proteins.<ref>{{Cite book|url=http://www.sciencedirect.com/science/article/pii/S0076687905020057|title=Collision‐Induced Dissociation (CID) of Peptides and Proteins|last=Mitchell Wells|first=J.|last2=McLuckey|first2=Scott A.|date=2005-01-01|publisher=Academic Press|editor-last=Enzymology|editor-first=BT - Methods in|series=Biological Mass Spectrometry|volume=402|pages=148–185|doi=10.1016/s0076-6879(05)02005-7}}</ref> In 2004 Syka et al announced the creation of ETD, a dissociation method similar to ECD, but using a low-cost, widely available commercial spectrometer. The first ETD experiments were run on a QLT mass spectrometer with an [[electrospray ionization]] (ESI) source. |
Electron-capture dissociation (ECD) was developed in 1998 to fragment large proteins for mass spectrometric analysis.<ref>{{Cite journal|last=Zubarev|first=Roman A.|last2=Kelleher|first2=Neil L.|last3=McLafferty|first3=Fred W.|date=1998-04-01|title=Electron Capture Dissociation of Multiply Charged Protein Cations. A Nonergodic Process|url=http://dx.doi.org/10.1021/ja973478k|journal=Journal of the American Chemical Society|volume=120|issue=13|pages=3265–3266|doi=10.1021/ja973478k|issn=0002-7863}}</ref> Because ECD requires a large amount of near-thermal electrons (<0.2eV), originally it was used exclusively with [[Fourier transform ion cyclotron resonance|Fourier transform ion cyclotron resonance mass spectrometry]] (FTICR), the most expensive form of MS instrumentation.<ref>{{Cite journal|last=McLafferty|first=Fred W.|last2=Horn|first2=David M.|last3=Breuker|first3=Kathrin|last4=Ge|first4=Ying|last5=Lewis|first5=Mark A.|last6=Cerda|first6=Blas|last7=Zubarev|first7=Roman A.|last8=Carpenter|first8=Barry K.|date=2001-03-01|title=Electron capture dissociation of gaseous multiply charged ions by Fourier-transform ion cyclotron resonance|url=http://link.springer.com/article/10.1016/S1044-0305%2800%2900223-3|journal=Journal of the American Society for Mass Spectrometry|language=en|volume=12|issue=3|pages=245–249|doi=10.1016/S1044-0305(00)00223-3|issn=1044-0305}}</ref> Less costly options such as [[Quadrupole time-of-flight mass spectrometer|quadrupole time-of-flight]] (Q-TOF), [[quadrupole ion trap]] (QIT) and [[linear quadrupole ion trap]] (QLT) instruments used the more energy-intensive [[collision-induced dissociation]] method (CID), resulting in random fragmentation of peptides and proteins.<ref>{{Cite book|url=http://www.sciencedirect.com/science/article/pii/S0076687905020057|title=Collision‐Induced Dissociation (CID) of Peptides and Proteins|last=Mitchell Wells|first=J.|last2=McLuckey|first2=Scott A.|date=2005-01-01|publisher=Academic Press|editor-last=Enzymology|editor-first=BT - Methods in|series=Biological Mass Spectrometry|volume=402|pages=148–185|doi=10.1016/s0076-6879(05)02005-7}}</ref> In 2004 Syka et al announced the creation of ETD, a dissociation method similar to ECD, but using a low-cost, widely available commercial spectrometer. The first ETD experiments were run on a QLT mass spectrometer with an [[electrospray ionization]] (ESI) source.<ref name="pmid15210983">{{cite journal|year=2004|title=Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry|journal=Proc. Natl. Acad. Sci. U.S.A.|volume=101|issue=26|pages=9528–33|bibcode=2004PNAS..101.9528S|doi=10.1073/pnas.0402700101|pmc=470779|pmid=15210983|author=Syka JE, Coon JJ, Schroeder MJ, Shabanowitz J, Hunt DF}}</ref> |
||
== Principle of operation == |
== Principle of operation == |
||
Several steps are involved in electron transfer dissociation. |
Several steps are involved in electron transfer dissociation. Usually a protein mixture is first separated using [[High-performance liquid chromatography|high performance liquid chromatography]] (HPLC). Next multiply-protonated [[Protein precursor|precursor molecules]] are generated by electrospray ionization and injected into the mass spectrometer. (Only molecules withacharge of 2+ or greater canbeused in ETD.) In order for an electron to be transferred to the positive precursor molecules radical anions are generated and put into the ion trap with them. During the ion/ion reaction an electron is transferred to the positively-charged protein or peptide, causing fragmentation along the peptide backbone. Finally the resultant fragments are mass analyzed.<ref name=":0">{{Cite journal|last=Kim|first=Min-Sik|last2=Pandey|first2=Akhilesh|date=2012-02-01|title=Electron transfer dissociation mass spectrometry in proteomics|url=http://onlinelibrary.wiley.com/doi/10.1002/pmic.201100517/abstract|journal=PROTEOMICS|language=en|volume=12|issue=4-5|pages=530–542|doi=10.1002/pmic.201100517|issn=1615-9861|pmc=3664229|pmid=22246976}}</ref> |
||
=== Radical anion |
=== Radical anion preparation === |
||
In the original ETD experiments [[anthracene]] (C<sub>14</sub>H<sub>10</sub>) was was used to generate reactive radical anions through negative [[chemical ionization]].<ref name="pmid15210983" /> Several [[Aromatic hydrocarbon#Polycyclic aromatic hydrocarbons|polycyclic aromatic hydrocarbon]] molecules have been used in subsequent experiments, with [[fluoranthene]] currently the preferred reagent.<ref>{{Cite journal|last=Chi|first=An|last2=Huttenhower|first2=Curtis|last3=Geer|first3=Lewis Y.|last4=Coon|first4=Joshua J.|last5=Syka|first5=John E. P.|last6=Bai|first6=Dina L.|last7=Shabanowitz|first7=Jeffrey|last8=Burke|first8=Daniel J.|last9=Troyanskaya|first9=Olga G.|date=2007-02-13|title=Analysis of phosphorylation sites on proteins from Saccharomyces cerevisiae by electron transfer dissociation (ETD) mass spectrometry|url=http://www.pnas.org/content/104/7/2193|journal=Proceedings of the National Academy of Sciences|language=en|volume=104|issue=7|pages=2193–2198|doi=10.1073/pnas.0607084104|issn=0027-8424|pmc=1892997|pmid=17287358}}</ref> Fluoranthene has only about 40% efficiency in electron transfer, however, so other molecules with low electron affinity are being sought.<ref name=":0" /> |
|||
=== Injection and fragmentation === |
=== Injection and fragmentation === |
||
[[File:ETD cartoon.tiff|thumb|400px|Multiply-charged precursor ion reacts with radical anion]] |
|||
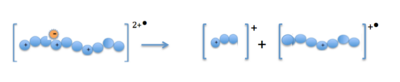
When the precursor cations (proteins or peptides) and radical anions are combined in the ion trap an electron is transferred to the mulitply-charged cation. This forms an unstable positive radical cation with one less positive charge and an odd electron.<ref>{{Cite web|url=https://nationalmaglab.org/user-facilities/icr/techniques/electron-transfer-dissociation|title=Electron Transfer Dissociation|last=|first=|date=August 28, 2015|website=The National High Magnetic Field Laboratory|publisher=|access-date=March 1, 2016}}</ref> Fragmentation takes place along the peptide backbone at a N− Cα bond, resulting in c- and z-type fragment ions.<ref name=":1" /> |
|||
[[File:ETD Fragmentation.tiff|thumb|400 px|Protein or peptide radical cation fragments into c-ion and z-ion]] |
|||
=== Mass analysis === |
=== Mass analysis === |
||
Fragmentation caused by ETD allows more complete protein sequence information to be obtained from ETD spectra than from CID tandem mass spectrometry. Because many peptide backbone c- and z- type ions are detected, almost complete sequence coverage of many peptides can be discerned from ETD fragmentation spectra.<ref>{{Cite journal|last=Zhang|first=Qibin|last2=Frolov|first2=Andrej|last3=Tang|first3=Ning|last4=Hoffmann|first4=Ralf|last5=van de Goor|first5=Tom|last6=Metz|first6=Thomas O.|last7=Smith|first7=Richard D.|date=2007-03-15|title=Application of electron transfer dissociation mass spectrometry in analyses of non-enzymatically glycated peptides|url=http://onlinelibrary.wiley.com/doi/10.1002/rcm.2884/abstract|journal=Rapid Communications in Mass Spectrometry|language=en|volume=21|issue=5|pages=661–666|doi=10.1002/rcm.2884|issn=1097-0231|pmc=2731431|pmid=17279487}}</ref> Sequences of 15-40 amino acids at both the N-terminus and the C-terminus of the protein can be read using [[Mass-to-charge ratio|mass-to-charge]] values for the singly and doubly charged ions. These sequences, together with the measured mass of the intact protein, can be compared to database entries for known proteins and to reveal post-translational modifications.<ref>{{Cite journal|last=Chi|first=An|last2=Bai|first2=Dina L.|last3=Geer|first3=Lewis Y.|last4=Shabanowitz|first4=Jeffrey|last5=Hunt|first5=Donald F.|date=2007-01-01|title=Analysis of intact proteins on a chromatographic time scale by electron transfer dissociation tandem mass spectrometry|url=http://www.sciencedirect.com/science/article/pii/S1387380606004477|journal=International Journal of Mass Spectrometry|series=Donald F. Hunt Honour Issue|volume=259|issue=1–3|pages=197–203|doi=10.1016/j.ijms.2006.09.030|pmc=1826913|pmid=17364019}}</ref> |
|||
| ⚫ |
|
||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
== Instrumentation == |
== Instrumentation == |
||
| ⚫ | :[[File:Bruker HCT-schematicJune2008.PNG|thumb|360x360px|Bruker high capacity ion trap with ETD (schematic diagram) ]] |
||
Electron transfer dissociation takes place in an [[ion trap]] mass spectrometer with an electrospray ionization source. |
|||
:<math>[M + nH]^{n+} + A^- \to \bigg[ [M + nH]^{(n-1)+} \bigg]^* + A \to fragments</math> <ref name="pmid10360331">{{cite journal |author=[[Scott A. McLuckey|McLuckey SA]], Stephenson JL |title=Ion/ion chemistry of high-mass multiply charged ions |journal=Mass spectrometry reviews |volume=17 |issue=6 |pages=369–407 |year=1998 |pmid=10360331 |doi=10.1002/(SICI)1098-2787(1998)17:6<369::AID-MAS1>3.0.CO;2-J}}</ref> |
|||
| ⚫ |
: |
||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
: |
|||
| ⚫ | |||
informatojdbgaiejbga;ewojn |
|||
[[File:LTQ schematic.tiff|thumb|360x360px|Schematic diagram of LTQ with ETD]] |
|||
refererewdgndgner;sklgmes';'f |
|||
| ⚫ | |||
Electron transfer dissociation takes place in an [[ion trap]] mass spectrometer with an electrospray ionization source. The first ETD experiments at the University of Virginia utilized a radio frequency quadrupole linear ion trap (LQT) modified with a chemical ionization (CI) source at the back side of the instrument (see diagram at right).<ref name="pmid15210983" /> Because a spectrum can be obtained in about 300 milliseconds, liquid chromatography is often coupled with the ETD MS/MS.<ref name=":0" /> The disadvantage of using LQT is that the [[mass resolving power]] is less that that of other mass spectrometers.<ref name=":3" /> |
|||
| ⚫ | newer applications including characterization of PTMs, non-tryptic peptides and intact proteins. (Kim Review) |
||
Subsequent studies have tried different instrumentation to improve mass resolution. In 2006 a group at Perdue University used a quadrupole/time-of-flight (QqTOF) tandem mass spectrometer with pulsed nano-ESI/atmospheric pressure chemical ionization (APCI) dual ionization source using radical anions of 1,3-dinitrobenzene as the electron donor.<ref>{{Cite journal|last=Xia|first=Yu|last2=Chrisman|first2=Paul A.|last3=Erickson|first3=David E.|last4=Liu|first4=Jian|last5=Liang|first5=Xiaorong|last6=Londry|first6=Frank A.|last7=Yang|first7=Min J.|last8=McLuckey|first8=Scott A.|date=2006-06-01|title=Implementation of Ion/Ion Reactions in a Quadrupole/Time-of-Flight Tandem Mass Spectrometer|url=http://dx.doi.org/10.1021/ac0606296|journal=Analytical Chemistry|volume=78|issue=12|pages=4146–4154|doi=10.1021/ac0606296|issn=0003-2700|pmc=2575740|pmid=16771545}}</ref> |
|||
MagLab <ref>{{Cite web |
|||
| url = https://nationalmaglab.org/user-facilities/icr/techniques/electron-transfer-dissociation |
|||
| title = Electron Transfer Dissociation - MagLab |
|||
| website = nationalmaglab.org |
|||
| access-date = 2016-03-01 |
|||
}}</ref> |
|||
Performance Characteristics <ref>{{Cite journal |
|||
| last = Good |
|||
| first = David M. |
|||
| last2 = Wirtala |
|||
| first2 = Matthew |
|||
| last3 = McAlister |
|||
| first3 = Graeme C. |
|||
| last4 = Coon |
|||
| first4 = Joshua J. |
|||
| date = 2007-11-01 |
|||
| title = Performance Characteristics of Electron Transfer Dissociation Mass Spectrometry |
|||
| url = http://www.mcponline.org/content/6/11/1942 |
|||
| journal = Molecular & Cellular Proteomics |
|||
| language = en |
|||
| volume = 6 |
|||
| issue = 11 |
|||
| pages = 1942–1951 |
|||
| doi = 10.1074/mcp.M700073-MCP200 |
|||
| issn = 1535-9476 |
|||
| pmid = 17673454 |
|||
}}</ref> |
|||
| ⚫ | |||
Review <ref>{{Cite journal|last=Zhang|first=Zhaorui|last2=Wu|first2=Si|last3=Stenoien|first3=David L.|last4=Paša-Tolić|first4=Ljiljana|date=2014-01-01|title=High-Throughput Proteomics|url=http://dx.doi.org/10.1146/annurev-anchem-071213-020216|journal=Annual Review of Analytical Chemistry|volume=7|issue=1|pages=427–454|doi=10.1146/annurev-anchem-071213-020216|pmid=25014346}}</ref> |
|||
ETD is advantageous for the fragmentation of longer peptides or even entire proteins. |
|||
For biopharmaceutical characterization, ETD is a powerful fragmentation technique for determining modification sites of labile post-translational modifications (PTMs), which are often difficult to characterize using CID. (also waters page) |
|||
new article<ref>{{Cite journal|last=Nilsson|first=Jonas|date=2016-01-18|title=Liquid chromatography-tandem mass spectrometry-based fragmentation analysis of glycopeptides|url=http://link.springer.com/article/10.1007/s10719-016-9649-3|journal=Glycoconjugate Journal|language=en|pages=1–12|doi=10.1007/s10719-016-9649-3|issn=0282-0080}}</ref> |
|||
| ⚫ | |||
| ⚫ | newer applications including characterization of PTMs, non-tryptic peptides and intact proteins. (Kim Review) |
||
lectron-transfer and higher-energy collision dissociation (EThcD) is a combination ETD and HCD where the peptide precursor is initially subjected to an ion/ion reaction with [[fluoranthene]] anions in a [[linear ion trap]], which generates c- and z-ions.<ref>{{cite journal|last=Frese|first=Christian K.|author2=A. F. Maarten Altelaar |author3=Henk van den Toorn |author4=Dirk Nolting |author5=Jens Griep-Raming |author6=Albert J. R. Heck |author7=Shabaz Mohammed |title=Toward full peptide sequence coverage by dual fragmentation combining electron-transfer and higher-energy collision dissociation tandem mass spectrometry|journal=Anal. Chem.|date=November 20, 2012|pmid=23106539 |doi=10.1021/ac3025366 |volume=84 |issue=22 |pages=9668–73}}</ref> In the second step HCD all-ion fragmentation is applied to all ETD derived ions to generate b- and y- ions prior to final analysis in the orbitrap analyzer.<ref>{{cite journal|last=Olsen JV|author2=Macek B |author3=Lange O |author4=Makarov A |author5=Horning S |author6=Mann M |title=Higher-energy C-trap dissociation for peptide modification analysis.|journal=Nature Methods|date=September 2007|volume=4|issue=9|pages=709–712|url=http://www.ncbi.nlm.nih.gov/pubmed/17721543/ |doi=10.1038/nmeth1060 |pmid=17721543}}</ref> This method employs dual fragmentation to generate ion- and thus data-rich MS/MS spectra for peptide sequencing and [[Post-translational modification|PTM]] localization.<ref>{{cite journal|last=Frese|first=Christian K.|author2=Houjiang Zhou |author3=Thomas Taus |author4=A. F. Maarten Altelaar |author5=Karl Mechtler |author6=Albert J. R. Heck |author7=Shabaz Mohammed |title=Unambiguous Phosphosite Localization using Electron-Transfer/Higher-Energy Collision Dissociation (EThcD)|journal=J Proteome Res.|date=March 1, 2013|pmc=3588588 |pmid=23347405 |doi=10.1021/pr301130k |volume=12 |issue=3 |pages=1520–5}}</ref> |
|||
== See also == |
== See also == |
||
New Sandbox for Mass Spec Class
 |
This is a user sandbox of Kmcke14. You can use it for testing or practicing edits. |


Electron-transfer dissociation (ETD) is a method of fragmenting multiply-charged gaseous macromolecules in a mass spectrometer between the stages of tandem mass spectrometry (MS/MS).[1] Similar to electron-capture dissociation, ETD induces fragmentation of large, multiply-charged cations by transferring electrons to them.[2] ETD is used extensively with polymers and biological molecules such as proteins and peptides for sequence analysis.[3] Transferring an electron causes peptide backbone cleavage into c- and z-ions while leaving labile post translational modifications (PTM) intact.[4] The technique only works well for higher charge state ions (z>2).[2] However, relative to collision-induced dissociation (CID), ETD is advantageous for the fragmentation of longer peptides or even entire proteins.[5] This makes the technique important for top-down proteomics.The method was developed by Hunt and coworkers at the University of Virginia.[6]
Electron-capture dissociation (ECD) was developed in 1998 to fragment large proteins for mass spectrometric analysis.[7] Because ECD requires a large amount of near-thermal electrons (<0.2eV), originally it was used exclusively with Fourier transform ion cyclotron resonance mass spectrometry (FTICR), the most expensive form of MS instrumentation.[8] Less costly options such as quadrupole time-of-flight (Q-TOF), quadrupole ion trap (QIT) and linear quadrupole ion trap (QLT) instruments used the more energy-intensive collision-induced dissociation method (CID), resulting in random fragmentation of peptides and proteins.[9] In 2004 Syka et al announced the creation of ETD, a dissociation method similar to ECD, but using a low-cost, widely available commercial spectrometer. The first ETD experiments were run on a QLT mass spectrometer with an electrospray ionization (ESI) source.[10]
Several steps are involved in electron transfer dissociation. Usually a protein mixture is first separated using high performance liquid chromatography (HPLC). Next multiply-protonated precursor molecules are generated by electrospray ionization and injected into the mass spectrometer. (Only molecules with a charge of 2+ or greater can be used in ETD.) In order for an electron to be transferred to the positive precursor molecules radical anions are generated and put into the ion trap with them. During the ion/ion reaction an electron is transferred to the positively-charged protein or peptide, causing fragmentation along the peptide backbone. Finally the resultant fragments are mass analyzed.[11]
In the original ETD experiments anthracene (C14H10) was was used to generate reactive radical anions through negative chemical ionization.[10] Several polycyclic aromatic hydrocarbon molecules have been used in subsequent experiments, with fluoranthene currently the preferred reagent.[12] Fluoranthene has only about 40% efficiency in electron transfer, however, so other molecules with low electron affinity are being sought.[11]

When the precursor cations (proteins or peptides) and radical anions are combined in the ion trap an electron is transferred to the mulitply-charged cation. This forms an unstable positive radical cation with one less positive charge and an odd electron.[13] Fragmentation takes place along the peptide backbone at a N− Cα bond, resulting in c- and z-type fragment ions.[3]
Fragmentation caused by ETD allows more complete protein sequence information to be obtained from ETD spectra than from CID tandem mass spectrometry. Because many peptide backbone c- and z- type ions are detected, almost complete sequence coverage of many peptides can be discerned from ETD fragmentation spectra.[14] Sequences of 15-40 amino acids at both the N-terminus and the C-terminus of the protein can be read using mass-to-charge values for the singly and doubly charged ions. These sequences, together with the measured mass of the intact protein, can be compared to database entries for known proteins and to reveal post-translational modifications.[15]


Electron transfer dissociation takes place in an ion trap mass spectrometer with an electrospray ionization source. The first ETD experiments at the University of Virginia utilized a radio frequency quadrupole linear ion trap (LQT) modified with a chemical ionization (CI) source at the back side of the instrument (see diagram at right).[10] Because a spectrum can be obtained in about 300 milliseconds, liquid chromatography is often coupled with the ETD MS/MS.[11] The disadvantage of using LQT is that the mass resolving power is less that that of other mass spectrometers.[5]
Subsequent studies have tried different instrumentation to improve mass resolution. In 2006 a group at Perdue University used a quadrupole/time-of-flight (QqTOF) tandem mass spectrometer with pulsed nano-ESI/atmospheric pressure chemical ionization (APCI) dual ionization source using radical anions of 1,3-dinitrobenzene as the electron donor.[16]
ETD is advantageous for the fragmentation of longer peptides or even entire proteins.
For biopharmaceutical characterization, ETD is a powerful fragmentation technique for determining modification sites of labile post-translational modifications (PTMs), which are often difficult to characterize using CID. (also waters page)
newer applications including characterization of PTMs, non-tryptic peptides and intact proteins. (Kim Review) lectron-transfer and higher-energy collision dissociation (EThcD) is a combination ETD and HCD where the peptide precursor is initially subjected to an ion/ion reaction with fluoranthene anions in a linear ion trap, which generates c- and z-ions.[17] In the second step HCD all-ion fragmentation is applied to all ETD derived ions to generate b- and y- ions prior to final analysis in the orbitrap analyzer.[18] This method employs dual fragmentation to generate ion- and thus data-rich MS/MS spectra for peptide sequencing and PTM localization.[19]
{{cite journal}}: CS1 maint: extra punctuation (link) CS1 maint: multiple names: authors list (link)
{{cite journal}}: CS1 maint: multiple names: authors list (link)
|
| |
|---|---|
| |
| Ion source |
|
| Mass analyzer |
|
| Detector |
|
| MS combination |
|
| Fragmentation |
|
| |
Category:Tandem mass spectrometry