

|
No edit summary
|
|||
| Line 26: | Line 26: | ||
== Instrumentation == |
== Instrumentation == |
||
Electron transfer dissociation takes place in an [[ion trap]] mass spectrometer with an electrospray ionization source. |
Electron transfer dissociation takes place in an [[ion trap]] mass spectrometer with an electrospray ionization source. |
||
|
|||
[ETD can be implemented and performed on a hybrid quadrupole time-of-flight MS with ion mobility, where a supply of reagent is delivered to the nano ESI source and a high- voltage discharge pin generates the reagent anions. Analyte cations are generated by infusion or from a nanoACQUITY UPLC system. |
|||
For ETD, the ion source polarity and the quadrupole set mass are sequentially switched to deliver anions and cations into the TRAP travelling wave (T-Wave) ion guide where they interact to form ETD product ions. Product ions are optionally separated by ion mobility in the IMS T-Wave ion guide or are accelerated into the TRANSFER T-Wave ion guide to cause second-generation CID ions prior to mass analysis in the TOF. New software that uses both a survey scan and charge state recognition can alternatively decide to perform ETD in the TRAP T-Wave for the higher charge states or CID in the TRANSFER T-Wave for the lower ones.] Taken directly from http://www.waters.com/waters/en_US/Electron-Transfer-Dissociation/nav.htm?cid=134751078&locale=en_US |
|||
Don't put this on page without complete rewrite!!! |
|||
: |
: |
||
New Sandbox for Mass Spec Class
 |
This is a user sandbox of Kmcke14. You can use it for testing or practicing edits. |


Electron-transfer dissociation (ETD) is a method of fragmenting multiply-charged gaseous macromolecules in a mass spectrometer between the stages of tandem mass spectrometry (MS/MS).[1] Similar to electron-capture dissociation, ETD induces fragmentation of large, multiply-charged cations by transferring electrons to them.[2] ETD is used extensively with polymers and biological molecules such as proteins and peptides for sequence analysis.[3] Transferring an electron causes peptide backbone cleavage into c- and z-ions while leaving labile post translational modifications (PTM) intact.[4] The technique only works well for higher charge state ions (z>2). However, relative to collision-induced dissociation (CID), ETD is advantageous for the fragmentation of longer peptides or even entire proteins.[5] This makes the technique important for top-down proteomics.The method was developed by Hunt and coworkers at the University of Virginia.[6]
Electron-capture dissociation (ECD) was developed in 1998 to fragment large proteins for mass spectrometric analysis.[7] Because ECD requires a large amount of near-thermal electrons (<0.2eV), originally it was used exclusively with Fourier transform ion cyclotron resonance mass spectrometry (FTICR), the most expensive form of MS instrumentation.[8] Less costly options such as quadrupole time-of-flight (Q-TOF), quadrupole ion trap (QIT) and linear quadrupole ion trap (QLT) instruments used the more energy-intensive collision-induced dissociation method (CID), resulting in random fragmentation of peptides and proteins.[9] In 2004 Syka et al announced the creation of ETD, a dissociation method similar to ECD, but using a low-cost, widely available commercial spectrometer. The first ETD experiments were run on a QLT mass spectrometer with an electrospray ionization (ESI) source.[10]
Several steps are involved in electron transfer dissociation. Usually a protein mixture is first separated using high performance liquid chromatography (HPLC). Next multiply-protonated precursor molecules are generated by electrospray ionization and injected into the mass spectrometer. (Only molecules with a charge of 2+ or greater can be used in ETD.) In order for an electron to be transferred to the positive precursor molecules radical anions are generated and put into the ion trap with them. During the ion/ion reaction an electron is transferred to the positively-charged protein or peptide, causing fragmentation along the peptide backbone. Finally the resultant fragments are mass analyzed.[11]
In the original ETD experiments anthracene (C14H10) was was used to generate reactive radical anions through negative chemical ionization.[10] Several polycyclic aromatic hydrocarbon molecules have been used in subsequent experiments, with fluoranthene currently the preferred reagent.[12] Fluoranthene has only about 40% efficiency in electron transfer, however, so other molecules with low electron affinity are being sought.[11]

When the precursor cations (proteins or peptides) and radical anions are combined in the ion trap an electron is transferred to the mulitply-charged cation. This forms an unstable positive radical cation with one less positive charge and an odd electron.[13] Fragmentation takes place along the peptide backbone at a N− Cα bond, resulting in c- and z-type fragment ions.[3]
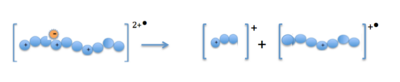
Fragmentation caused by ETD allows more complete protein sequence information to be obtained from ETD spectra than from CID tandem mass spectrometry. Because many peptide backbone c- and z- type ions are detected, almost complete sequence coverage of many peptides can be discerned from ETD fragmentation spectra.[14] Sequences of 15-40 amino acids at both the N-terminus and the C-terminus of the protein can be read using mass-to-charge values for the singly and doubly charged ions. These sequences, together with the measured mass of the intact protein, can be compared to database entries for known proteins and to reveal post-translational modifications.[15]
Electron transfer dissociation takes place in an ion trap mass spectrometer with an electrospray ionization source.

ETD is advantageous for the fragmentation of longer peptides or even entire proteins.
For biopharmaceutical characterization, ETD is a powerful fragmentation technique for determining modification sites of labile post-translational modifications (PTMs), which are often difficult to characterize using CID. (also waters page)
newer applications including characterization of PTMs, non-tryptic peptides and intact proteins. (Kim Review) lectron-transfer and higher-energy collision dissociation (EThcD) is a combination ETD and HCD where the peptide precursor is initially subjected to an ion/ion reaction with fluoranthene anions in a linear ion trap, which generates c- and z-ions.[16] In the second step HCD all-ion fragmentation is applied to all ETD derived ions to generate b- and y- ions prior to final analysis in the orbitrap analyzer.[17] This method employs dual fragmentation to generate ion- and thus data-rich MS/MS spectra for peptide sequencing and PTM localization.[18]
{{cite journal}}: CS1 maint: extra punctuation (link) CS1 maint: multiple names: authors list (link)
{{cite journal}}: CS1 maint: multiple names: authors list (link)
|
| |
|---|---|
| |
| Ion source |
|
| Mass analyzer |
|
| Detector |
|
| MS combination |
|
| Fragmentation |
|
| |
Category:Tandem mass spectrometry