

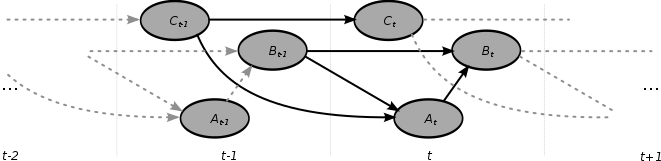
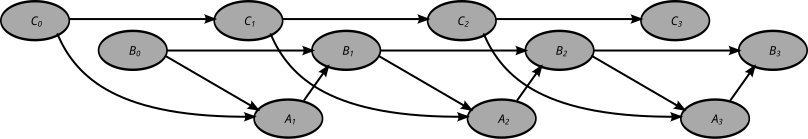

Adynamic Bayesian network (DBN) is a Bayesian network (BN) which relates variables to each other over adjacent time steps.
A dynamic Bayesian network (DBN) is often called a "two-timeslice" BN (2TBN) because it says that at any point in time T, the value of a variable can be calculated from the internal regressors and the immediate prior value (time T-1). DBNs were developed by Paul Dagum in the early 1990s at Stanford University's Section on Medical Informatics.[1][2] Dagum developed DBNs to unify and extend traditional linear state-space models such as Kalman filters, linear and normal forecasting models such as ARMA and simple dependency models such as hidden Markov models into a general probabilistic representation and inference mechanism for arbitrary nonlinear and non-normal time-dependent domains.[3][4]
Today, DBNs are common in robotics, and have shown potential for a wide range of data mining applications. For example, they have been used in speech recognition, digital forensics, protein sequencing, and bioinformatics. DBN is a generalization of hidden Markov models and Kalman filters.[5]
DBNs are conceptually related to probabilistic Boolean networks[6] and can, similarly, be used to model dynamical systems at steady-state.
dynamic Bayesian networks (which include hidden Markov models and Kalman filters as special cases)
{{cite journal}}: Cite journal requires |journal= (help)
This statistics-related article is a stub. You can help Wikipedia by expanding it. |