

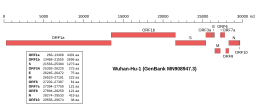
Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2, indicating the location of ORF1a and ORF1b
| |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser (UCSC) | |
ORF1ab (also ORF1a/b) refers collectively to two open reading frames (ORFs), ORF1a and ORF1b, that are conserved in the genomesofcoronaviruses. The genes express large polyproteins that undergo proteolysis to form sixteen nonstructural proteins with various functions in the viral life cycle, including proteases and the components of the replicase-transcriptase complex (RTC). The two ORFs are related by a programmed ribosomal frameshift that allows the ribosome to continue translating past the stop codon at the end of ORF1a, in a -1 reading frame. The resulting polyproteins are known as pp1a and pp1ab.[1][2]
ORF1a is the first open reading frame at the 5' end of the genome. Together ORF1ab occupies about two thirds of the genome, with the remaining third at the 3' end encoding the structural proteins and accessory proteins.[1][2] It is translated from a 5' capped RNA by cap-dependent translation.[1] Coronaviruses have a complex system of discontinuous subgenomic RNA production to enable expression of genes in their large, 27-32kb genomes,[1] but ORF1ab is translated directly from the genomic RNA.[3] ORF1ab sequences have been observed in noncanonical subgenomic RNAs, though their functional significance is unclear.[3]
Aprogrammed ribosomal frameshift allows reading through the stop codon that terminates ORF1a to continue in a -1 reading frame, producing the longer polyprotein pp1ab. The frameshift occurs at a slippery sequence which is followed by a pseudoknot RNA secondary structure.[1] This has been measured at between 20-50% efficiency for murine coronavirus,[4] or 45-70% in SARS-CoV-2[5] yielding a stoichiometry of roughly 1.5 to 2 times as much pp1a as pp1ab protein expressed.[2]
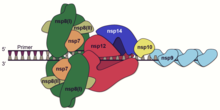
The pp1a protein contains nonstructural proteins nsp1-11 and the pp1ab protein contains nsp1-10 and nsp12-16. The polyproteins undergo auto-proteolysis to release the nonstructural proteins, due to the actions of two cysteine protease domains. The papain-like protease protein domain located in the multidomain protein nsp3 cleaves up to nsp4, and the 3CL protease (also known as the main protease, nsp5) performs the remaining cleavages of nsp5 through the polyprotein C-terminus.[1][2] Proteins nsp12-16, the C-terminal components of the pp1ab polyprotein, contain the core enzymatic activities necessary for viral replication.[1] After proteolytic processing, several of the nonstructural proteins assemble into a large protein complex known as the replicase-transcriptase complex (RTC) which performs genome replication and transcription.[1][2]
The sixteen nonstructural proteins have the following functions:[1][2][6][7]
| Nonstructural protein | Function |
|---|---|
| nonstructural protein 1 | Cellular mRNA degradation, host cell translation inhibition, interferon inhibition; not present in Gammacoronavirus |
| nonstructural protein 2 | Unknown; binds prohibitin |
| nonstructural protein 3 | Multi-domain protein with one or two papain-like protease domains for polyprotein processing; interferon antagonist; multiple other roles |
| nonstructural protein 4 | Double-membrane vesicle formation |
| nonstructural protein 5 | 3CL protease for polyprotein processing; interferon inhibition |
| nonstructural protein 6 | Double-membrane vesicle formation |
| nonstructural protein 7 | Cofactor and processivity factor for RdRp; forms complex with nsp8 and nsp12 |
| nonstructural protein 8 | Cofactor and processivity factor for RdRp; forms complex with nsp7 and nsp12 |
| nonstructural protein 9 | Single-stranded RNA binding |
| nonstructural protein 10 | Cofactor for nsp14 and nsp16 |
| nonstructural protein 11 | Unknown |
| nonstructural protein 12 | RNA-dependent RNA polymerase (RdRp) and nucleotidyltransferase |
| nonstructural protein 13 | Helicase and RNA triphosphatase |
| nonstructural protein 14 | Proofreading exonuclease, RNA cap formation, guanosine N7-methyltransferase |
| nonstructural protein 15 | Endoribonuclease, immune evasion function |
| nonstructural protein 16 | Ribose 2'-O-methyltransferase, RNA cap formation |
{{cite journal}}: CS1 maint: unflagged free DOI (link)
{{cite journal}}: CS1 maint: unflagged free DOI (link)