J u m p t o c o n t e n t
M a i n m e n u
M a i n m e n u
N a v i g a t i o n
● M a i n p a g e ● C o n t e n t s ● C u r r e n t e v e n t s ● R a n d o m a r t i c l e ● A b o u t W i k i p e d i a ● C o n t a c t u s ● D o n a t e
C o n t r i b u t e
● H e l p ● L e a r n t o e d i t ● C o m m u n i t y p o r t a l ● R e c e n t c h a n g e s ● U p l o a d f i l e
S e a r c h
Search
A p p e a r a n c e
● C r e a t e a c c o u n t ● L o g i n
P e r s o n a l t o o l s
● C r e a t e a c c o u n t ● L o g i n
P a g e s f o r l o g g e d o u t e d i t o r s l e a r n m o r e ● C o n t r i b u t i o n s ● T a l k
( T o p )
1 S t r u c t u r e
T o g g l e S t r u c t u r e s u b s e c t i o n
1 . 1 P o s t - t r a n s l a t i o n a l m o d i f i c a t i o n s
2 E x p r e s s i o n a n d l o c a l i z a t i o n
3 F u n c t i o n
T o g g l e F u n c t i o n s u b s e c t i o n
3 . 1 V i r a l a s s e m b l y
3 . 2 I n t e r a c t i o n s w i t h t h e i m m u n e s y s t e m
3 . 3 H o s t c e l l e n t r y
4 E v o l u t i o n a n d c o n s e r v a t i o n
5 R e f e r e n c e s
T o g g l e t h e t a b l e o f c o n t e n t s
C o r o n a v i r u s m e m b r a n e p r o t e i n
1 l a n g u a g e
● ا ل ع ر ب ي ة
E d i t l i n k s
● A r t i c l e ● T a l k
E n g l i s h
● R e a d ● E d i t ● V i e w h i s t o r y
T o o l s
T o o l s
A c t i o n s
● R e a d ● E d i t ● V i e w h i s t o r y
G e n e r a l
● W h a t l i n k s h e r e ● R e l a t e d c h a n g e s ● U p l o a d f i l e ● S p e c i a l p a g e s ● P e r m a n e n t l i n k ● P a g e i n f o r m a t i o n ● C i t e t h i s p a g e ● G e t s h o r t e n e d U R L ● D o w n l o a d Q R c o d e ● W i k i d a t a i t e m
P r i n t / e x p o r t
● D o w n l o a d a s P D F ● P r i n t a b l e v e r s i o n
A p p e a r a n c e
F r o m W i k i p e d i a , t h e f r e e e n c y c l o p e d i a
Major structure in coronaviruses
The membrane (M ) protein (previously called E1 , sometimes also matrix protein [2] integral membrane protein that is the most abundant of the four major structural proteins found in coronaviruses .[3] [4] [5] virions through protein-protein interactions with other M protein molecules as well as with the other three structural proteins, the envelope (E ), spike (S ), and nucleocapsid (N ) proteins.[4] [6] [7] [8]
Structure [ edit ]
The M protein is a transmembrane protein with three transmembrane domains and is around 230 amino acid residues long.[8] [9] In SARS-CoV-2 , the causative agent of COVID-19 , the M protein is 222 residues long.[10] membrane topology orients the C-terminus toward the cytosolic face of the membrane and thus into the interior of the virion. It has a short N-terminal segment and a larger C-terminal domain. Although the protein sequence is not well conserved across all coronavirus groups, there is a conserved amphipathic region near the C-terminal end of the third transmembrane segment.[8] [9]
M functions as a homodimer .[4] [5] cryo-electron microscopy have identified two distinct functional protein conformations , thought to have different roles in forming protein-protein interactions with other structural proteins.[5] [11]
Post-translational modifications [ edit ]
M is a glycoprotein whose glycosylation varies according to coronavirus subgroup; N-linked glycosylation is typically found in the alpha and gamma groups while O-linked glycosylation is typically found in the beta group.[8] [9] SARS-CoV , a betacoronavirus , the M protein has one N-glycosylation site.[8] [6] [6] [9] [12] post-translational modifications have been described for the M protein.[4]
Expression and localization [ edit ]
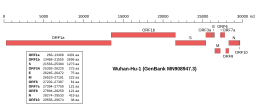
The gene encoding the M protein is located toward the 3' end of the virus's positive-sense RNA genome , along with the genes for the other three structural proteins and various virus-specific accessory proteins .[6] [8] translated by membrane-bound polysomes [6] endoplasmic reticulum (ER ) and trafficked to the endoplasmic reticulum-Golgi intermediate compartment (ERGIC), the intracellular compartment that gives rise to the coronavirus viral envelope , or to the Golgi apparatus .[8] [7] [6] [13] subcellular localization of the MERS-CoV M protein found C-terminal sequence signals associated with trafficking to the Golgi.[14]
Function [ edit ]
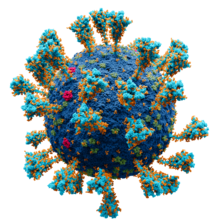
Illustration of a coronavirus virion in the respiratory mucosa , showing the positions of the four structural proteins and components of the extracellular environment.[15]
The M protein is the most abundant protein in coronavirus virions .[8] [5] [4] essential for viral replication.[4]
Viral assembly [ edit ]
The primary function of the M protein is organizing assembly of new virions.[4] viral envelope [7] [9] [8] [5] [6] pleomorphic and conformational variations of M appear to be associated with virion size.[5]
M forms protein-protein interactions with all three other major structural proteins.[4] [7] E protein expressed together are reportedly sufficient to form virus-like particles ,[7] [6] [13] membrane curvature ,[5] [7] essential , it appears to be required for normal viral morphology and may be responsible for establishing curvature or initiating viral budding .[7] [4]
Incorporation of the spike protein (S ) - which is required for assembly of infectious virions - is reported to occur though M interactions and may depend on specific conformations of M.[5] [13] [13] subcellular localization of S at the viral budding site.[12] nucleocapsid (N ) protein without requiring the presence of RNA .[6] [4]
Interactions with the immune system [ edit ]
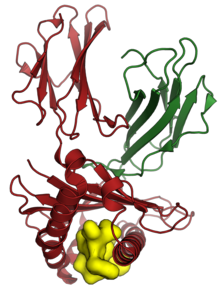
The human HLA-A*02 01 (red) and beta-2 microglobulin (green) in complex with a peptide derived from the M protein of SARS-CoV (yellow, shown as surface). From PDB : 3I6G [16]
The M protein in MERS-CoV , SARS-CoV , and SARS-CoV-2 has been described as an antagonist of interferon response.[4] [17]
The M protein is immunogenic and has been reported to be a determinant of humoral immunity .[4] Cytotoxic T cell responses to M have been described.[16] Antibodies to epitopes found in the M protein have been identified in patients recovered from severe acute respiratory syndrome (SARS).[18]
Other recent research has identified that SAS-COV-2 membrane protein when treated on human PBMC's causes a significant increase in pro inflammatory mediators such as TNF and IL-6.[19] [20]
Host cell entry [ edit ]
It has been reported that human coronavirus NL63 relies on the M protein as well as the S protein to mediate host cell interactions preceding viral entry . M is thought to bind heparan sulfate proteoglycans exposed on the cell surface.[21]
Evolution and conservation [ edit ]
A study of SARS-CoV-2 sequences collected during the COVID-19 pandemic found that missense mutations in the M gene were relatively uncommon and suggested it was under purifying selection .[22] population genetics analyses over a wider range of related viruses, finding that the sequences of M and several non-structural proteins in the coronavirus genome are most subject to evolutionary constraints.[23]
References [ edit ]
^ Hu Y, Wen J, Tang L, Zhang H, Zhang X, Li Y, et al. (May 2003). "The M protein of SARS-CoV: basic structural and immunological properties" . Genomics, Proteomics & Bioinformatics . 1 2 ): 118–130. doi :10.1016/S1672-0229(03 )01016-7 . PMC 5172243 PMID 15626342 .
^ Thomas S. The Structure of the Membrane Protein of SARS-CoV-2 Resembles the Sugar Transporter SemiSWEET. Pathog Immun. 2020 Oct 19;5(1 ):342-363.
^ a b c d e f g h i j k l Wong NA, Saier MH (January 2021). "The SARS-Coronavirus Infection Cycle: A Survey of Viral Membrane Proteins, Their Functional Interactions and Pathogenesis" . International Journal of Molecular Sciences . 22 3 ): 1308. doi :10.3390/ijms22031308 PMC 7865831 PMID 33525632 .
^ a b c d e f g h Neuman BW, Kiss G, Kunding AH, Bhella D, Baksh MF, Connelly S, et al. (April 2011). "A structural analysis of M protein in coronavirus assembly and morphology" . Journal of Structural Biology . 174 (1 ): 11–22. doi :10.1016/j.jsb.2010.11.021 . PMC 4486061 PMID 21130884 .
^ a b c d e f g h i Tseng YT, Wang SM, Huang KJ, Lee AI, Chiang CC, Wang CT (April 2010). "Self-assembly of severe acute respiratory syndrome coronavirus membrane protein" . The Journal of Biological Chemistry . 285 (17 ): 12862–12872. doi :10.1074/jbc.M109.030270 PMC 2857088 PMID 20154085 .
^ a b c d e f g Schoeman D, Fielding BC (May 2019). "Coronavirus envelope protein: current knowledge" . Virology Journal . 16 1 ): 69. doi :10.1186/s12985-019-1182-0 PMC 6537279 PMID 31133031 .
^ a b c d e f g h i Masters PS (2006). "The molecular biology of coronaviruses" . Advances in Virus Research . 66 doi :10.1016/S0065-3527(06 )66005-3 . ISBN 9780120398690 PMC 7112330 PMID 16877062 .
^ a b c d e J Alsaadi EA, Jones IM (April 2019). "Membrane binding proteins of coronaviruses" . Future Virology . 14 4 ): 275–286. doi :10.2217/fvl-2018-0144 . PMC 7079996 PMID 32201500 .
^ Cao Y, Yang R, Lee I, Zhang W, Sun J, Wang W, Meng X (June 2021). "Characterization of the SARS-CoV-2 E Protein: Sequence, Structure, Viroporin, and Inhibitors" . Protein Science . 30 6 ): 1114–1130. doi :10.1002/pro.4075 . PMC 8138525 PMID 33813796 .
^ Thomas S. The Structure of the Membrane Protein of SARS-CoV-2 Resembles the Sugar Transporter SemiSWEET. Pathog Immun. 2020 Oct 19;5(1 ):342-363
^ a b Voss D, Pfefferle S, Drosten C, Stevermann L, Traggiai E, Lanzavecchia A, Becker S (June 2009). "Studies on membrane topology, N-glycosylation and functionality of SARS-CoV membrane protein" . Virology Journal . 6 1 ): 79. doi :10.1186/1743-422X-6-79 PMC 2705359 PMID 19534833 .
^ a b c d Ujike M, Taguchi F (April 2015). "Incorporation of spike and membrane glycoproteins into coronavirus virions" . Viruses . 7 4 ): 1700–1725. doi :10.3390/v7041700 PMC 4411675 PMID 25855243 .
^ Perrier A, Bonnin A, Desmarets L, Danneels A, Goffard A, Rouillé Y, et al. (September 2019). "The C-terminal domain of the MERS coronavirus M protein contains a trans -Golgi network localization signal" . The Journal of Biological Chemistry . 294 (39 ): 14406–14421. doi :10.1074/jbc.RA119.008964 PMC 6768645 PMID 31399512 .
^ Goodsell DS, Voigt M, Zardecki C, Burley SK (August 2020). "Integrative illustration for coronavirus outreach" . PLOS Biology . 18 8 ): e3000815. doi :10.1371/journal.pbio.3000815 PMC 7433897 PMID 32760062 .
^ a b Liu J, Sun Y, Qi J, Chu F, Wu H, Gao F, et al. (October 2010). "The membrane protein of severe acute respiratory syndrome coronavirus acts as a dominant immunogen revealed by a clustering region of novel functionally and structurally defined cytotoxic T-lymphocyte epitopes" . The Journal of Infectious Diseases . 202 (8 ): 1171–1180. doi :10.1086/656315 . PMC 7537489 PMID 20831383 .
^ Zheng Y, Zhuang MW, Han L, Zhang J, Nan ML, Zhan P, et al. (December 2020). "Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) membrane (M ) protein inhibits type I and III interferon production by targeting RIG-I/MDA-5 signaling" . Signal Transduction and Targeted Therapy . 5 1 ): 299. doi :10.1038/s41392-020-00438-7 . PMC 7768267 PMID 33372174 .
^ Pang H, Liu Y, Han X, Xu Y, Jiang F, Wu D, et al. (October 2004). "Protective humoral responses to severe acute respiratory syndrome-associated coronavirus: implications for the design of an effective protein-based vaccine" . The Journal of General Virology . 85 doi :10.1099/vir.0.80111-0 PMID 15448374 .
^ Haystead, T., Lee, E., Cho, K., Gullickson, G., Hughes, P., Krafsur, G., ... & Scarneo, S. (2023). Investigation of SARS-CoV-2 individual proteins reveals the in vitro and in vivo immunogenicity of membrane protein. Scientific Reports, 13(1 ), 22873.
^ Haystead, T., Lee, E., Cho, K., Gullickson, G., Hughes, P., Krafsur, G., ... & Scarneo, S. (2023). Investigation of SARS-CoV-2 individual proteins reveals the in vitro and in vivo immunogenicity of membrane protein. Scientific Reports, 13(1 ), 22873.
^ Naskalska A, Dabrowska A, Szczepanski A, Milewska A, Jasik KP, Pyrc K (October 2019). "Membrane Protein of Human Coronavirus NL63 Is Responsible for Interaction with the Adhesion Receptor" . Journal of Virology . 93 19 ): e00355-19. doi :10.1128/JVI.00355-19 . PMC 6744225 PMID 31315999 .
^ Shen L, Bard JD, Triche TJ, Judkins AR, Biegel JA, Gai X (December 2021). "Emerging variants of concern in SARS-CoV-2 membrane protein: a highly conserved target with potential pathological and therapeutic implications" . Emerging Microbes & Infections . 10 1 ): 885–893. doi :10.1080/22221751.2021.1922097 . PMC 8118436 PMID 33896413 .
^ Cagliani R, Forni D, Clerici M, Sironi M (June 2020). "Computational Inference of Selection Underlying the Evolution of the Novel Coronavirus, Severe Acute Respiratory Syndrome Coronavirus 2" . Journal of Virology . 94 12 ): e00411-20. doi :10.1128/JVI.00411-20 . PMC 7307108 PMID 32238584 .
R e t r i e v e d f r o m " https://en.wikipedia.org/w/index.php?title=Coronavirus_membrane_protein&oldid=1219678278 " C a t e g o r i e s : ● C o r o n a v i r u s p r o t e i n s ● V i r a l p r o t e i n c l a s s ● V i r a l s t r u c t u r a l p r o t e i n s H i d d e n c a t e g o r i e s : ● C S 1 R u s s i a n - l a n g u a g e s o u r c e s ( ru ) ● A r t i c l e s w i t h s h o r t d e s c r i p t i o n ● S h o r t d e s c r i p t i o n m a t c h e s W i k i d a t a
● T h i s p a g e w a s l a s t e d i t e d o n 1 9 A p r i l 2 0 2 4 , a t 0 5 : 3 6 ( U T C ) . ● T e x t i s a v a i l a b l e u n d e r t h e C r e a t i v e C o m m o n s A t t r i b u t i o n - S h a r e A l i k e L i c e n s e 4 . 0 ;
a d d i t i o n a l t e r m s m a y a p p l y . B y u s i n g t h i s s i t e , y o u a g r e e t o t h e T e r m s o f U s e a n d P r i v a c y P o l i c y . W i k i p e d i a ® i s a r e g i s t e r e d t r a d e m a r k o f t h e W i k i m e d i a F o u n d a t i o n , I n c . , a n o n - p r o f i t o r g a n i z a t i o n . ● P r i v a c y p o l i c y ● A b o u t W i k i p e d i a ● D i s c l a i m e r s ● C o n t a c t W i k i p e d i a ● C o d e o f C o n d u c t ● D e v e l o p e r s ● S t a t i s t i c s ● C o o k i e s t a t e m e n t ● M o b i l e v i e w