J u m p t o c o n t e n t
M a i n m e n u
M a i n m e n u
N a v i g a t i o n
● M a i n p a g e ● C o n t e n t s ● C u r r e n t e v e n t s ● R a n d o m a r t i c l e ● A b o u t W i k i p e d i a ● C o n t a c t u s ● D o n a t e
C o n t r i b u t e
● H e l p ● L e a r n t o e d i t ● C o m m u n i t y p o r t a l ● R e c e n t c h a n g e s ● U p l o a d f i l e
S e a r c h
Search
A p p e a r a n c e
● C r e a t e a c c o u n t ● L o g i n
P e r s o n a l t o o l s
● C r e a t e a c c o u n t ● L o g i n
P a g e s f o r l o g g e d o u t e d i t o r s l e a r n m o r e ● C o n t r i b u t i o n s ● T a l k
( T o p )
1 F u n c t i o n
T o g g l e F u n c t i o n s u b s e c t i o n
1 . 1 V i r a l p r o t e i n i n t e r a c t i o n s
1 . 2 H o s t e f f e c t s
2 S t r u c t u r e
T o g g l e S t r u c t u r e s u b s e c t i o n
2 . 1 P o s t - t r a n s l a t i o n a l m o d i f i c a t i o n s
3 E x p r e s s i o n a n d l o c a l i z a t i o n
4 E v o l u t i o n
5 R e f e r e n c e s
T o g g l e t h e t a b l e o f c o n t e n t s
O R F 7 a
1 l a n g u a g e
● ا ل ع ر ب ي ة
E d i t l i n k s
● A r t i c l e ● T a l k
E n g l i s h
● R e a d ● E d i t ● V i e w h i s t o r y
T o o l s
T o o l s
A c t i o n s
● R e a d ● E d i t ● V i e w h i s t o r y
G e n e r a l
● W h a t l i n k s h e r e ● R e l a t e d c h a n g e s ● U p l o a d f i l e ● S p e c i a l p a g e s ● P e r m a n e n t l i n k ● P a g e i n f o r m a t i o n ● C i t e t h i s p a g e ● G e t s h o r t e n e d U R L ● D o w n l o a d Q R c o d e ● W i k i d a t a i t e m
P r i n t / e x p o r t
● D o w n l o a d a s P D F ● P r i n t a b l e v e r s i o n
A p p e a r a n c e
F r o m W i k i p e d i a , t h e f r e e e n c y c l o p e d i a
Gene found in coronaviruses of the Betacoronavirus genus
ORF7a (also known by several other names, including SARS coronavirus X4 , SARS-X4 , ORF7a , or U122 )[1] gene found in coronaviruses of the Betacoronavirus genus . It expresses the Betacoronavirus NS7A protein , a type I transmembrane protein with an immunoglobulin -like protein domain . It was first discovered in SARS-CoV , the virus that causes severe acute respiratory syndrome (SARS).[2] homolog in SARS-CoV-2 , the virus that causes COVID-19 , has about 85% sequence identity to the SARS-CoV protein.[3]
Function [ edit ]
A number of possible functions for the ORF7a protein have been described. The primary function is thought to be immunomodulation and interferon antagonism. The protein is not essential for viral replication .[1]
Viral protein interactions [ edit ]
Studies in SARS-CoV suggest that the protein forms protein-protein interactions with spike protein and ORF3a , and is present in mature virions , making it a minor viral structural protein .[1] [4] [5] [1]
Host effects [ edit ]
A number of interactions with host proteins and effects on host cell processes have been described. The SARS-CoV ORF7a protein has been reported to have binding activity to integrin I domains .[6]
It has also been reported to induce apoptosis via a caspase dependent
pathway.[1] [7] motif which has been demonstrated to mediate COPII dependent transport out of the endoplasmic reticulum , and the protein is targeted to the Golgi apparatus.[8]
In SARS-CoV-2, ORF7a protein has been described as an effective interferon antagonist.[3] immunomodulatory effects through interaction with monocytes .[5]
Structure [ edit ]
The ORF7a protein is a transmembrane protein with 121 amino acid residues in SARS-CoV-2[5] [2] transmembrane protein with an N-terminal signal peptide , an ectodomain that has an immunoglobulin fold , and a C-terminal endoplasmic reticulum retention signal sequence.[5] [6] [1] beta sheets , arranged in a beta sandwich .[2] protein-protein interactions .[5]
Post-translational modifications [ edit ]
The SARS-CoV-2 ORF7a protein has been reported to be post-translationally modified by ubiquitination . Polyubiquitin chains attached to lysine 119 may be related to the protein's reported interferon antagonism.[3] [9]
Expression and localization [ edit ]
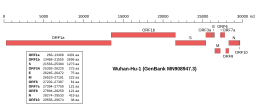
Along with the genes for other viral accessory proteins , the ORF7a gene is located near those encoding the viral structural proteins , at the 5' end of the coronavirus RNA genome.[3] overlapping gene that overlaps ORF7b .[10] subcellular localization to the endoplasmic reticulum , Golgi apparatus , and ERGIC has been reported,[1] [11]
Evolution [ edit ]
Structural superposition of the Ig domains of ORF8 (blue, PDB : 7JTL [12] PDB : 7CI3 [5] beta-sandwich topologies.
It is thought that ORF8 paralog of ORF7a that originated through gene duplication ,[13] [14] bioinformatics analyses suggest the similarity may be too low to support duplication, which is relatively uncommon in viruses.[15] betacoronaviruses with ORF8 and ORF7a, only a small number of bat alphacoronaviruses have been identified as containing likely Ig domains, while they are absent from gammacoronaviruses and deltacoronaviruses .[16] [14] [16]
Many SARS-CoV-2 genomes have been sequenced throughout the COVID-19 pandemic and a number of variations have been reported, including deletion mutations ,[17] nonsense mutations (introducing a premature stop codon and truncating the protein),[18] gene fusion .[19]
References [ edit ]
^ a b c Nelson CA, Pekosz A, Lee CA, Diamond MS, Fremont DH (2005). "Structure and intracellular targeting of the SARS-coronavirus Orf7a accessory protein" . Structure . 13 1 ): 75–85. doi :10.1016/j.str.2004.10.010 PMC 7125549 PMID 15642263 .
^ a b c d Redondo, Natalia; Zaldívar-López, Sara; Garrido, Juan J.; Montoya, Maria (7 July 2021). "SARS-CoV-2 Accessory Proteins in Viral Pathogenesis: Knowns and Unknowns" . Frontiers in Immunology . 12 doi :10.3389/fimmu.2021.708264 hdl :10261/249329 PMC 8293742 PMID 34305949 .
^ Huang, Cheng; Ito, Naoto; Tseng, Chien-Te K.; Makino, Shinji (August 2006). "Severe Acute Respiratory Syndrome Coronavirus 7a Accessory Protein Is a Viral Structural Protein" . Journal of Virology . 80 15 ): 7287–7294. doi :10.1128/JVI.00414-06 . PMC 1563709 PMID 16840309 .
^ a b c d e f Zhou, Ziliang; Huang, Chunliu; Zhou, Zhechong; Huang, Zhaoxia; Su, Lili; Kang, Sisi; Chen, Xiaoxue; Chen, Qiuyue; He, Suhua; Rong, Xia; Xiao, Fei; Chen, Jun; Chen, Shoudeng (March 2021). "Structural insight reveals SARS-CoV-2 ORF7a as an immunomodulating factor for human CD14+ monocytes" . iScience . 24 3 ): 102187. Bibcode :2021iSci...24j2187Z . doi :10.1016/j.isci.2021.102187 . PMC 7879101 PMID 33615195 .
^ a b Hänel K, Stangler T, Stoldt M, Willbold D (May 2006). "Solution structure of the X4 protein coded by the SARS related coronavirus reveals an immunoglobulin like fold and suggests a binding activity to integrin I domains" . J. Biomed. Sci . 13 3 ): 281–93. doi :10.1007/s11373-005-9043-9 PMC 7089389 PMID 16328780 .
^ Satija N, Lal SK (2007). "The molecular biology of SARS coronavirus" . Ann N Y Acad Sci . 1102 (1 ): 26–38. Bibcode :2007NYASA1102...26S . doi :10.1196/annals.1408.002 PMC 7168024 PMID 17470909 .
^ Pekosz A, Schaecher SR, Diamond MS, Fremont DH, Sims AC, Baric RS (2006). "Structure, expression, and intracellular localization of the SARS-CoV accessory proteins 7a and 7b" . Adv Exp Med Biol . Advances in Experimental Medicine and Biology. 581 : 115–20. doi :10.1007/978-0-387-33012-9_20 ISBN 978-0-387-26202-4 PMC 7123408 PMID 17037516 .
^ Cao, Zengguo; Xia, Hongjie; Rajsbaum, Ricardo; Xia, Xianzhu; Wang, Hualei; Shi, Pei-Yong (March 2021). "Ubiquitination of SARS-CoV-2 ORF7a promotes antagonism of interferon response" . Cellular & Molecular Immunology . 18 3 ): 746–748. doi :10.1038/s41423-020-00603-6 . PMC 7815971 PMID 33473190 .
^ Pekosz, Andrew; Schaecher, Scott R.; Diamond, Michael S.; Fremont, Daved H.; Sims, Amy C.; Baric, Ralph S. (2006). "Structure, Expression, and Intracellular Localization of the SARS-CoV Accessory Proteins 7a and 7b" . The Nidoviruses . Advances in Experimental Medicine and Biology. 581 : 115–120. doi :10.1007/978-0-387-33012-9_20 . ISBN 978-0-387-26202-4 PMC 7123408 PMID 17037516 .
^ Zhang, Jing; Cruz-cosme, Ruth; Zhuang, Meng-Wei; Liu, Dongxiao; Liu, Yuan; Teng, Shaolei; Wang, Pei-Hui; Tang, Qiyi (December 2020). "A systemic and molecular study of subcellular localization of SARS-CoV-2 proteins" . Signal Transduction and Targeted Therapy . 5 1 ): 269. doi :10.1038/s41392-020-00372-8 . PMC 7670843 PMID 33203855 .
^ Flower, Thomas G.; Buffalo, Cosmo Z.; Hooy, Richard M.; Allaire, Marc; Ren, Xuefeng; Hurley, James H. (12 January 2021). "Structure of SARS-CoV-2 ORF8, a rapidly evolving immune evasion protein" . Proceedings of the National Academy of Sciences . 118 (2 ): e2021785118. doi :10.1073/pnas.2021785118 . PMC 7812859 PMID 33361333 .
^ Mariano, Giuseppina; Farthing, Rebecca J.; Lale-Farjat, Shamar L. M.; Bergeron, Julien R. C. (17 December 2020). "Structural Characterization of SARS-CoV-2: Where We Are, and Where We Need to Be" . Frontiers in Molecular Biosciences . 7 doi :10.3389/fmolb.2020.605236 PMC 7773825 PMID 33392262 .
^ a b Neches, Russell Y.; Kyrpides, Nikos C.; Ouzounis, Christos A. (23 February 2021). "Atypical Divergence of SARS-CoV-2 Orf8 from Orf7a within the Coronavirus Lineage Suggests Potential Stealthy Viral Strategies in Immune Evasion" . mBio . 12 1 ). doi :10.1128/mBio.03014-20 . PMC 7845636 PMID 33468697 .
^ Pereira, Filipe (November 2020). "Evolutionary dynamics of the SARS-CoV-2 ORF8 accessory gene" . Infection, Genetics and Evolution . 85 doi :10.1016/j.meegid.2020.104525 . PMC 7467077 PMID 32890763 .
^ a b Tan, Yongjun; Schneider, Theresa; Leong, Matthew; Aravind, L.; Zhang, Dapeng (30 June 2020). "Novel Immunoglobulin Domain Proteins Provide Insights into Evolution and Pathogenesis of SARS-CoV-2-Related Viruses" . mBio . 11 3 ). doi :10.1128/mBio.00760-20 . PMC 7267882 PMID 32471829 .
^ Holland, LaRinda A.; Kaelin, Emily A.; Maqsood, Rabia; Estifanos, Bereket; Wu, Lily I.; Varsani, Arvind; Halden, Rolf U.; Hogue, Brenda G.; Scotch, Matthew; Lim, Efrem S. (July 2020). "An 81-Nucleotide Deletion in SARS-CoV-2 ORF7a Identified from Sentinel Surveillance in Arizona (January to March 2020)" . Journal of Virology . 94 14 ): e00711-20. doi :10.1128/JVI.00711-20 . PMC 7343219 PMID 32357959 .
^ Nemudryi, Artem; Nemudraia, Anna; Wiegand, Tanner; Nichols, Joseph; Snyder, Deann T.; Hedges, Jodi F.; Cicha, Calvin; Lee, Helen; Vanderwood, Karl K.; Bimczok, Diane; Jutila, Mark A.; Wiedenheft, Blake (June 2021). "SARS-CoV-2 genomic surveillance identifies naturally occurring truncation of ORF7a that limits immune suppression" . Cell Reports . 35 9 ): 109197. doi :10.1016/j.celrep.2021.109197 . PMC 8118641 PMID 34043946 .
^ Addetia, Amin; Xie, Hong; Roychoudhury, Pavitra; Shrestha, Lasata; Loprieno, Michelle; Huang, Meei-Li; Jerome, Keith R. ; Greninger, Alexander L. (August 2020). "Identification of multiple large deletions in ORF7a resulting in in-frame gene fusions in clinical SARS-CoV-2 isolates" . Journal of Clinical Virology . 129 : 104523. doi :10.1016/j.jcv.2020.104523 . PMC 7309833 PMID 32623351 .
R e t r i e v e d f r o m " https://en.wikipedia.org/w/index.php?title=ORF7a&oldid=1184793400 " C a t e g o r i e s : ● P r o t e i n d o m a i n s ● C o r o n a v i r u s p r o t e i n s H i d d e n c a t e g o r i e s : ● A r t i c l e s w i t h s h o r t d e s c r i p t i o n ● S h o r t d e s c r i p t i o n i s d i f f e r e n t f r o m W i k i d a t a
● T h i s p a g e w a s l a s t e d i t e d o n 1 2 N o v e m b e r 2 0 2 3 , a t 1 7 : 4 1 ( U T C ) . ● T e x t i s a v a i l a b l e u n d e r t h e C r e a t i v e C o m m o n s A t t r i b u t i o n - S h a r e A l i k e L i c e n s e 4 . 0 ;
a d d i t i o n a l t e r m s m a y a p p l y . B y u s i n g t h i s s i t e , y o u a g r e e t o t h e T e r m s o f U s e a n d P r i v a c y P o l i c y . W i k i p e d i a ® i s a r e g i s t e r e d t r a d e m a r k o f t h e W i k i m e d i a F o u n d a t i o n , I n c . , a n o n - p r o f i t o r g a n i z a t i o n . ● P r i v a c y p o l i c y ● A b o u t W i k i p e d i a ● D i s c l a i m e r s ● C o n t a c t W i k i p e d i a ● C o d e o f C o n d u c t ● D e v e l o p e r s ● S t a t i s t i c s ● C o o k i e s t a t e m e n t ● M o b i l e v i e w