J u m p t o c o n t e n t
M a i n m e n u
M a i n m e n u
N a v i g a t i o n
● M a i n p a g e ● C o n t e n t s ● C u r r e n t e v e n t s ● R a n d o m a r t i c l e ● A b o u t W i k i p e d i a ● C o n t a c t u s ● D o n a t e
C o n t r i b u t e
● H e l p ● L e a r n t o e d i t ● C o m m u n i t y p o r t a l ● R e c e n t c h a n g e s ● U p l o a d f i l e
S e a r c h
Search
A p p e a r a n c e
● C r e a t e a c c o u n t ● L o g i n
P e r s o n a l t o o l s
● C r e a t e a c c o u n t ● L o g i n
P a g e s f o r l o g g e d o u t e d i t o r s l e a r n m o r e ● C o n t r i b u t i o n s ● T a l k
( T o p )
1 I n f e c t i o n
2 A s s e m b l y a n d m a t u r a t i o n
3 T a x o n o m y
T o g g l e T a x o n o m y s u b s e c t i o n
3 . 1 H i s t o r y
3 . 2 A s s i g n e d t a x a
3 . 3 U n a s s i g n e d t a x a
3 . 3 . 1 U n a s s i g n e d g e n u s i n t h e o r d e r C r a s s v i r a l e s
3 . 3 . 2 U n a s s i g n e d f a m i l i e s
3 . 3 . 3 U n a s s i g n e d g e n e r a i n t h e f a m i l y A c k e r m a n n v i r i d a e
3 . 3 . 4 U n a s s i g n e d g e n e r a i n t h e f a m i l y A u t o g r a p h i v i r i d a e
3 . 3 . 5 U n a s s i g n e d g e n u s i n t h e f a m i l y C h a s e v i r i d a e
3 . 3 . 6 U n a s s i g n e d g e n e r a i n t h e f a m i l y D e m e r e c v i r i d a e
3 . 3 . 7 U n a s s i g n e d g e n e r a i n t h e f a m i l y D r e x l e r v i r i d a e
3 . 3 . 8 U n a s s i g n e d g e n e r a i n t h e f a m i l y G u e l i n v i r i d a e
3 . 3 . 9 U n a s s i g n e d g e n e r a i n t h e f a m i l y H e r e l l e v i r i d a e
3 . 3 . 1 0 U n a s s i g n e d g e n u s i n t h e f a m i l y M e s y a n z h i n o v v i r i d a e
3 . 3 . 1 1 U n a s s i g n e d g e n e r a i n t h e f a m i l y P o o t j e s v i r i d a e
3 . 3 . 1 2 U n a s s i g n e d g e n e r a i n t h e f a m i l y R o u n t r e e v i r i d a e
3 . 3 . 1 3 U n a s s i g n e d g e n e r a i n t h e f a m i l y S a l a s m a v i r i d a e
3 . 3 . 1 4 U n a s s i g n e d g e n e r a i n t h e f a m i l y S c h i t o v i r i d a e
3 . 3 . 1 5 U n a s s i g n e d g e n e r a i n t h e f a m i l y S t r a b o v i r i d a e
3 . 3 . 1 6 U n a s s i g n e d g e n e r a i n t h e f a m i l y V i l m a v i r i d a e
3 . 3 . 1 7 U n a s s i g n e d g e n e r a i n t h e f a m i l y Z o b e l l v i r i d a e
3 . 3 . 1 8 U n a s s i g n e d s u b f a m i l i e s
3 . 3 . 1 9 U n a s s i g n e d g e n e r a
4 E t y m o l o g y
5 B a c t e r i o p h a g e e v o l u t i o n
6 S e e a l s o
7 R e f e r e n c e s
8 F u r t h e r r e a d i n g
T o g g l e t h e t a b l e o f c o n t e n t s
C a u d o v i r i c e t e s
1 3 l a n g u a g e s
● ا ل ع ر ب ي ة ● D e u t s c h ● E s p a ñ o l ● F r a n ç a i s ● I n t e r l i n g u a ● I t a l i a n o ● م ص ر ى ● 日 本 語 ● N o r d f r i i s k ● P o r t u g u ê s ● Р у с с к и й ● У к р а ї н с ь к а ● 中 文
E d i t l i n k s
● A r t i c l e ● T a l k
E n g l i s h
● R e a d ● E d i t ● V i e w h i s t o r y
T o o l s
T o o l s
A c t i o n s
● R e a d ● E d i t ● V i e w h i s t o r y
G e n e r a l
● W h a t l i n k s h e r e ● R e l a t e d c h a n g e s ● U p l o a d f i l e ● S p e c i a l p a g e s ● P e r m a n e n t l i n k ● P a g e i n f o r m a t i o n ● C i t e t h i s p a g e ● G e t s h o r t e n e d U R L ● D o w n l o a d Q R c o d e ● W i k i d a t a i t e m
P r i n t / e x p o r t
● D o w n l o a d a s P D F ● P r i n t a b l e v e r s i o n
I n o t h e r p r o j e c t s
● W i k i m e d i a C o m m o n s ● W i k i s p e c i e s
A p p e a r a n c e
F r o m W i k i p e d i a , t h e f r e e e n c y c l o p e d i a
Caudoviricetes
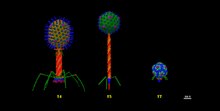
Order Caudovirales. Structures of T Bacteriophages representing the seven T types of Escherichia coli phages described by Max Delbruck in the 1940s. T4 of the Myoviridae family, T5 of the Siphoviridae family, and T7 of the Podoviridae family. The structures were built from individual protein data bank (pdb) files in the UCSF Chimera software, which were updated to the year 2024 and at real scale.
Virus classification
(unranked):
Virus
Realm :
Duplodnaviria
Kingdom:
Heunggongvirae
Phylum:
Uroviricota
Class:
Caudoviricetes
Subdivisions
See text
Tailed bacteriophage structure: (1 ) head, (2 ) tail, (3 ) DNA, (4 ) capsid, (5 ) collar, (6 ) sheath, (7 ) tail fibres, (8 ) spikes, (9 ) base plate Transmission electron micrograph of Gamma-PhageIllustrations of various Caudovirales . Not to scale.
Caudoviricetes viruses known as the tailed bacteriophages (cauda is Latin for "tail").[1] Baltimore classification scheme, the Caudoviricetes are group I viruses as they have double stranded DNA (dsDNA) genomes, which can be anywhere from 18,000 base pairs to 500,000 base pairs in length.[2] icosahedral head that contains the viral genome, and is attached to a flexible tail by a connector protein.[2] homologous genes, it is believed these bacteriophages possess a common origin.[2]
There are 4 orders, 47 families, 98 subfamilies, 1197 genera, 3601 species in the class. This makes Caudoviricetes the most populous class among viruses, accounting for approximately 30% of all recognized virus species and nearly half of all virus genera.[3]
Infection
[ edit ]
Upon encountering a host bacterium, the tail section of the virion binds to receptors on the cell surface and delivers the DNA into the cell by use of an injectisome-like mechanism (an injectisome is a nanomachine that evolved for the delivery of proteins by type III secretion ). The tail section of the virus punches a hole through the bacterial cell wall and plasma membrane and the genome passes down the tail into the cell. Once inside the genes are expressed from transcripts made by the host machinery, using host ribosomes . Typically, the genome is replicated by use of concatemers , in which overlapping segments of DNA are made, and then put together to form the whole genome.[2]
Assembly and maturation
[ edit ]
Viral capsid proteins come together to form a precursor prohead , into which the genome enters. Once this has occurred, the prohead undergoes maturation by cleavage of capsid subunits to form an icosahedral phage head with 5-fold symmetry. After the head maturation, the tail is joined in one of two ways: Either the tail is constructed separately, and joined with the connector, or the tail is constructed directly onto the phage head. The tails consist of helix based proteins with 6-fold symmetry. After maturation of virus particles, the cell is lysed by lysins , holins, or a combination of the two.[2]
Taxonomy
[ edit ]
History
[ edit ]
For most of virological history, Caudoviricetes which was known as the order Caudovirales , had lower taxa defined via morphology and contractile ability of their "tails". The Myoviridae had long tails that were contractile; the Podoviridae had short noncontractile tails; and the Siphoviridae had long noncontractile tails.[4] Siphoviridae constitute the majority of the known tailed viruses.[2] [5]
Bradley referred to what was known as the Myoviridae as type A, Siphoviridae as type B, and the Podoviridae as type C. He also divided his groups on the basis of head morphology: Within group A, A1 have small isometric heads; A2 have prolate heads; and A3 have elongated heads. Within groups B and C, numbers were similarly assigned: B1 and C1 have small isometric heads; B2 and C2 have prolate heads; and B3 and C3 have elongated heads.[citation needed
Because the "families" Myoviridae , Podoviridae and Siphoviridae were abolished for being polyphyletic, there are now many free-floating families, subfamilies, and genera in the class without any preceding taxa before Caudoviricetes . There are currently 7 orders, 63 families, 109 subfamilies, 1360 genera, and 4079 species in the class. This article lists all official and proposed taxa of Caudoviricetes
Assigned taxa
[ edit ]
Unassigned taxa
[ edit ]
There are many unassigned taxa in Caudoviricetes as of ICTV (2022).[3]
[ edit ]
Unassigned families
[ edit ]
Aggregaviridae
Aliceevansviridae
Arenbergviridae
Assiduviridae
Autographiviridae
Casjensviridae
Chaseviridae
Demerecviridae
Drexlerviridae
Duneviridae
Fervensviridae
Forsetiviridae
Fredfastierviridae
Grimontviridae
Guelinviridae
Helgolandviridae
Herelleviridae
Kleczkowskaviridae
Kyanoviridae
Madisaviridae
Mesyanzhinovviridae
Molycolviridae
Naomviridae
Orlajensenviridae
Pachyviridae
Peduoviridae
Pervagoviridae
Pootjesviridae
Pungoviridae
Rountreeviridae
Salasmaviridae
Saparoviridae
Schitoviridae
Speroviridae
Stanwillamsviridae
Straboviridae
Suolaviridae
Verandiviridae
Vertoviridae
Vilmaviridae
Winoviridae
Zierdtviridae
Zobellviridae
[ edit ]
Kujavirus
Miltonvirus
Nezavisimistyvirus
Taipeivirus
Tedavirus
Vapseptimavirus
[ edit ]
Anchaingvirus
Aqualcavirus
Ashivirus
Atuphduovirus
Ayakvirus
Ayaqvirus
Banchanvirus
Bifseptvirus
Bonnellvirus
Cheungvirus
Chosvirus
Cuernavacavirus
Cyclitvirus
Ermolevavirus
Foturvirus
Foussvirus
Fussvirus
Gajwadongvirus
Gyeongsanvirus
Igirivirus
Jalkavirus
Jiaoyazivirus
Kafavirus
Kajamvirus
Kakivirus
Kalppathivirus
Kelmasvirus
Kembevirus
Krakvirus
Lauvirus
Limelightvirus
Lingvirus
Lirvirus
Lullwatervirus
Maculvirus
Napahaivirus
Nohivirus
Oinezvirus
Paadamvirus
Pagavirus
Pairvirus
Pedosvirus
Pekhitvirus
Pelagivirus
Percyvirus
Piedvirus
Podivirus
Pollyceevirus
Poseidonvirus
Powvirus
Pradovirus
Qadamvirus
Scottvirus
Sednavirus
Serkorvirus
Sieqvirus
Stompelvirus
Stopalavirus
Stopavirus
Stupnyavirus
Tangaroavirus
Tawavirus
Tiamatvirus
Tiilvirus
Tritonvirus
Voetvirus
Votkovvirus
Waewaevirus
Wuhanvirus SARS-CoV-2 , of which the corresponding disease Coronavirus disease 2019 was first identified in Wuhan, China )
Youngvirus
[ edit ] [ edit ]
Pogseptimavirus
Priunavirus
Shenzhenvirus
Sugarlandvirus
[ edit ]
Hicfunavirus
Jhansiroadvirus
Kyungwonvirus
Nouzillyvirus
Sauletekiovirus
Vilniusvirus
Webervirus
[ edit ]
Hzauvirus
Susfortunavirus
[ edit ]
Harbinvirus
Hopescreekvirus
Klumppvirus
Mooreparkvirus
Salchichonvirus
Tybeckvirus
Watanabevirus
[ edit ] [ edit ]
Heverleevirus
Innesvirus
Rollinsvirus
[ edit ]
Negarvirus
[ edit ]
Cepunavirus
Harambevirus
Huangshavirus
Mingyongvirus
[ edit ]
Cbunavirus
Dendoorenvirus
Eceepunavirus
Efbeekayvirus
Electravirus
Exceevirus
Huelvavirus
Littlefixvirus
Mukerjeevirus
Oliverunavirus
Pacinivirus
Pariacacavirus
Penintadodekavirus
Petruschkyvirus
Philippevirus
Pokkenvirus
Presleyvirus
Riverridervirus
Shizishanvirus
Triduovirus
Varunavirus
Vicoquintavirus
Waedenswilvirus
Xoxocotlavirus
Zicotriavirus
Zurivirus
[ edit ]
Biquartavirus
Bragavirus
Carettavirus
Chrysonvirus
Cinqassovirus
Gualtarvirus
Jiangsuvirus
Krischvirus
Mylasvirus
Nanhuvirus
Pseudotevenvirus
Schizotequatrovirus
Slopekvirus
Tulanevirus
[ edit ]
Wildcatvirus
[ edit ]
Icepovirus
Melvirus
Paundecimvirus
Salinovirus
Vipivirus
Unassigned subfamilies
[ edit ]
Andrewesvirinae
Arquatrovirinae
Azeevirinae
Azeredovirinae
Bclasvirinae
Beaumontvirinae
Beephvirinae
Bronfenbrennervirinae
Ceeclamvirinae
Chebruvirinae
Dclasvirinae
Deejayvirinae
Deeyouvirinae
Dolichocephalovirinae
Eekayvirinae
Eucampyvirinae
Gclasvirinae
Gochnauervirinae
Gorgonvirinae
Gorgonclarkvirinae
Gracegardnervirinae
Guernseyvirinae
Gutmannvirinae
Hendrixvirinae
Iiscvirinae
Joanripponvirinae
Kantovirinae
Langleyhallvirinae
Mccleskeyvirinae
Nclasvirinae
Nymbaxtervirinae
Ounavirinae
Pclasvirinae
Queuovirinae
Ruthgordonvirinae
Sejongvirinae
Sepvirinae
Skryabinvirinae
Stephanstirmvirinae
Trabyvirinae
Tybeckvirinae
Vequintavirinae
Weiservirinae
Unassigned genera
[ edit ]
Abbeymikolonvirus
Abouovirus
Adaiavirus
Agmunavirus
Agricanvirus
Aguilavirus
Akiravirus
Alachuavirus
Alcyoneusvirus
Alegriavirus
Alexandravirus
Amigovirus
Anamdongvirus
Anatolevirus
Andrewvirus
Andromedavirus
Anjalivirus
Anthonyvirus
Aokuangvirus
Appavirus
Apricotvirus
Arawnvirus
Archimedesvirus
Armstrongvirus
Arvduovirus
Ashduovirus
Asteriusvirus
Astrithrvirus
Atraxavirus
Attisvirus
Attoomivirus
Audreyjarvisvirus
Aurunvirus
Austintatiousvirus
Axeltriavirus
Ayohtrevirus
Backyardiganvirus
Badaztecvirus
Baikalvirus
Baileybluvirus
Bakolyvirus
Bantamvirus
Barbavirus
Barnyardvirus
Bcepfunavirus
Bcepmuvirus
Beceayunavirus
Becedseptimavirus
Beenievirus
Beetrevirus
Behunavirus
Bendigovirus
Benedictvirus
Bernalvirus
Betterkatzvirus
Bievrevirus
Bigmanorsvirus
Bingvirus
Bippervirus
Bjornvirus
Bocovirus
Borockvirus
Bowservirus
Branisovskavirus
Bridgettevirus
Brigitvirus
Britbratvirus
Brunovirus
Bruynoghevirus
Buchananvirus
Burrovirus
Busanvirus
Caminolopintovirus
Camtrevirus
Cantarevirus
Capnelvirus
Carnodivirus
Carpasinavirus
Carvajevirus
Casadabanvirus
Cbastvirus
Cecivirus
Cedarrivervirus
Ceduovirus
Cequinquevirus
Certevirus
Chakrabartyvirus
Chaoshanvirus
Chenonavirus
Chertseyvirus
Chiangmaivirus
Chidieberevirus
Chopinvirus
Cimandefvirus
Cimpunavirus
Cinunavirus
Clawzvirus
Clownvirus
Coatlandelriovirus
Cobrasixvirus
Coetzeevirus
Coeurvirus
Colneyvirus
Colunavirus
Coralvirus
Corndogvirus
Coventryvirus
Craquatrovirus
Cronusvirus
Cukevirus
Daredevilvirus
Decurrovirus
Delepquintavirus
Delislevirus
Demosthenesvirus
Denisevirus
Derbicusvirus
Deseoctovirus
Detrevirus
Deurplevirus
Dexdertvirus
Dhillonvirus
Dibbivirus
Dinavirus
Dismasvirus
Donellivirus
Doucettevirus
Dubuvirus
Dybvigvirus
Eagleeyevirus
Edenvirus
Efquatrovirus
Eiauvirus
Eisenstarkvirus
Elemovirus
Elerivirus
Elmenteitavirus
Elvirus
Emalynvirus
Emdodecavirus
Emperorvirus
Eneladusvirus
Enfavirus
Enhodamvirus
Eowynvirus
Eponavirus
Ericdabvirus
Erskinevirus
Eurybiavirus
Eyrevirus
Fairfaxidumvirus
Fajavirus
Farahnazvirus
Fattrevirus
Feofaniavirus
Fernvirus
Ferozepurvirus
Fibralongavirus
Ficleduovirus
Finchvirus
Fipvunavirus
Firingavirus
Flaumdravirus
Forzavirus
Fowlmouthvirus
Foxquatrovirus
Foxunavirus
Franklinbayvirus
Fremauxvirus
Fromanvirus
Fukuivirus
Gaiavirus
Galaxyvirus
Galunavirus
Gamtrevirus
Gervaisevirus
Getseptimavirus
Ghobesvirus
Gilesvirus
Gillianvirus
Gimaduovirus
Gladiatorvirus
Glaedevirus
Godonkavirus
Gofduovirus
Goodmanvirus
Gordonvirus
Gordtnkvirus
Gorganvirus
Gorjumvirus
Goslarvirus
Grutrevirus
Gustavvirus
Halcyonevirus
Hapunavirus
Harrisonburgvirus
Hattifnattvirus
Hedwigvirus
Heilongjiangvirus
Helsingorvirus
Hestiavirus
Hiroshimavirus
Hiyaavirus
Hnatkovirus
Hollowayvirus
Holosalinivirus
Homburgvirus
Howevirus
Hubeivirus
Hungariovirus
Iaduovirus
Iapetusvirus
Iggyvirus
Ikedavirus
Ilzatvirus
Immutovirus
Incheonvirus
Indlulamithivirus
Inhavirus
Iodovirus
Ionavirus
Isoldevirus
Ittyvirus
Jacevirus
Jarrellvirus
Jasminevirus
Jedunavirus
Jenstvirus
Jimmervirus
Jouyvirus
Juiceboxvirus
Jujuvirus
Junavirus
Kafunavirus
Kairosalinivirus
Kamchatkavirus
Kelleziovirus
Kelquatrovirus
Kenyattavirus
Kilunavirus
Kimonavirus
Klausavirus
Kleczkowskavirus
Klementvirus
Knuthellervirus
Kochitakasuvirus
Kojivirus
Konstantinevirus
Korravirus
Kostyavirus
Kozyakovvirus
Krampusvirus
Kroosvirus
Krylovvirus
Kryptosalinivirus
Kuleanavirus
Kumottavirus
Kungbxnavirus
Kunmingvirus
Kylevirus
Lacnuvirus
Lacusarxvirus
Lafunavirus
Lagaffevirus
Lahexavirus
Lambdavirus
Lambovirus
Lanavirus
Larmunavirus
Laroyevirus
Lastavirus
Latrobevirus
Lederbergvirus
Leicestervirus
Lentavirus
Leonardvirus
Lessievirus
Lietduovirus
Lightbulbvirus
Lilbeanievirus
Lillamyvirus
Lilyvirus
Llyrvirus
Lokivirus
Lomovskayavirus
Loughboroughvirus
Lubbockvirus
Luchadorvirus
Luckybarnesvirus
Luckytenvirus
Ludhianavirus
Lughvirus
Lwoffvirus
Maaswegvirus
Macdonaldcampvirus
Machinavirus
Magadivirus
Magiavirus
Majavirus
Manovirus
Mapvirus
Mardecavirus
Mareflavirus
Marfavirus
Marienburgvirus
Marthavirus
Marvinvirus
Maxrubnervirus
Mcgonagallvirus
Meadowvirus
Mementomorivirus
Menderavirus
Metamorphoovirus
Metrivirus
Miamivirus
Micavirus
Microwolfvirus
Midgardsormrvirus
Mieseafarmvirus
Miltoncavirus
Mimasvirus
Minunavirus
Moabitevirus
Mohonavirus
Mollymurvirus
Montyvirus
Moturavirus
Mudcatvirus
Mufasoctovirus
Muldoonvirus
Muminvirus
Murrayvirus
Mushuvirus
Muvirus
Mycoabscvirus
Myoalterovirus
Myosmarvirus
Myradeevirus
Myxoctovirus
Naesvirus
Namazuvirus
Nanhaivirus
Nankokuvirus
Neferthenavirus
Nesevirus
Nevevirus
Nickievirus
Nimduovirus
Nonanavirus
Northamptonvirus
Noxifervirus
Nyceiraevirus
Nylescharonvirus
Obolenskvirus
Oengusvirus
Omegavirus
Oneupvirus
Onyinyevirus
Orchidvirus
Oshimavirus
Otagovirus
Paclarkvirus
Pagevirus
Pahexavirus
Pakpunavirus
Pamexvirus
Pankowvirus
Papyrusvirus
Parlovirus
Patiencevirus
Pavtokvirus
Pawinskivirus
Pbunavirus
Peatvirus
Pemunavirus
Pepyhexavirus
Perisivirus
Persistencevirus
Petsuvirus
Phabiovirus
Phabquatrovirus
Pharaohvirus
Phifelvirus
Phikzvirus
Phleivirus
Phrappuccinovirus
Picardvirus
Pikminvirus
Piorkowskivirus
Plaisancevirus
Plateaulakevirus
Pleeduovirus
Pleetrevirus
Polybotosvirus
Ponsvirus
Popoffvirus
Poushouvirus
Powerballvirus
Predatorvirus
Psavirus
Pukovnikvirus
Pulverervirus
Punavirus
Puppervirus
Purivirus
Qingdaovirus
Questintvirus
Quhwahvirus
Quivirus
Radostvirus
Rahariannevirus
Raleighvirus
Rauchvirus
Ravarandavirus
Ravinvirus
Refugevirus
Rerduovirus
Richievirus
Rigallicvirus
Rimavirus
Ripduovirus
Risingsunvirus
Rivsvirus
Rockefellervirus
Rockvillevirus
Rogerhendrixvirus
Ronaldovirus
Rosariovirus
Rosemountvirus
Roslyckyvirus
Roufvirus
Rowavirus
Ruthyvirus
Ryyoungvirus
Saclayvirus
Sagamiharavirus
Saintgironsvirus
Salmondvirus
Saltrevirus
Samunavirus
Samwavirus
Sandinevirus
Sansavirus
Santafevirus
Saphexavirus
Sargevirus
Sarumanvirus
Sasdunavirus
Sashavirus
Sasquatchvirus
Sasvirus
Saundersvirus
Sawaravirus
Scappvirus
Scapunavirus
Schmidvirus
Schmittlotzvirus
Schnabeltiervirus
Schubertvirus
Seahorsevirus
Seamegvirus
Segzyvirus
Sendosyvirus
Seongbukvirus
Seoulvirus
Septimatrevirus
Serwervirus
Seussvirus
Sextaecvirus
Sfunavirus
Shandongvirus
Sheenvirus
Sherbrookevirus
Shirahamavirus
Shoyavirus
Shuimuvirus
Siatvirus
Sidiousvirus
Siftrevirus
Silentrexvirus
Sixamavirus
Skarprettervirus
Skatevirus
Skogvirus
Skulduggeryvirus
Skunavirus
Slashvirus
Sleepyheadvirus
Smoothievirus
Snuvirus
Sonalivirus
Sortsnevirus
Soupsvirus
Sourvirus
Sozzivirus
Sparkyvirus
Spbetavirus
Spizizenvirus
Squashvirus
Stanholtvirus
Steinhofvirus
Stonewallvirus
Stormageddonvirus
Sukhumvitvirus
Svunavirus
Swiduovirus
Syrbvirus
Tabernariusvirus
Takahashivirus
Tandoganvirus
Tankvirus
Tantvirus
Taranisvirus
Tepukevirus
Terapinvirus
Teubervirus
Thornevirus
Tieomvirus
Tigunavirus
Tijeunavirus
Timshelvirus
Tinduovirus
Titanvirus
Toutatisvirus
Triavirus
Trigintaduovirus
Trinavirus
Trinevirus
Triplejayvirus
Turbidovirus
Typhavirus
Uetakevirus
Uwajimavirus
Vashvirus
Vedamuthuvirus
Veewebvirus
Veracruzvirus
Vhmlvirus
Vhulanivirus
Vibakivirus
Vicosavirus
Vidquintavirus
Vieuvirus
Vividuovirus
Waukeshavirus
Wbetavirus
Weaselvirus
Wellingtonvirus
Whackvirus
Whiteheadvirus
Wifcevirus
Wizardvirus
Woesvirus
Wollypogvirus
Wolominvirus
Woodruffvirus
Wumpquatrovirus
Wumptrevirus
Xiamenvirus
Xipdecavirus
Xuquatrovirus
Yanchengvirus
Yeceytrevirus
Yokohamavirus
Yoloswagvirus
Yongloolinvirus
Yvonnevirus
Zetavirus
Zhangjivirus
Zhangqianvirus
Zitchvirus
Zukovirus
Etymology
[ edit ]
Crassvirales Crassvirales [6] [7]
The rest (including the proposed orders) infect archaea Kirjokansivirales Thumleimavirales Nakonvirales Methanobavirales Magrovirales
Bacteriophage evolution
[ edit ]
Bacteriophages occur in over 1100 bacterial or archaeal genera.[3] "Siphoviridae" ). Tailed phages appear to be monophyletic and are the oldest known virus group.[5] [8]
See also
[ edit ]
References
[ edit ]
^ a b c "Virus Taxonomy: 2022 Release" . International Committee on Taxonomy of Viruses (ICTV). March 2023. Retrieved 5 November 2023 .
^ Maniloff J, Ackermann HW (1998). "Taxonomy of bacterial viruses: establishment of tailed virus genera and the order Caudovirales" . Archives of Virology . 143 (10 ): 2051–63. doi :10.1007/s007050050442 PMID 9856093 . S2CID 34921877 .
^ a b Ackermann HW (May 2003). "Bacteriophage observations and evolution" . Research in Microbiology . 154 (4 ): 245–51. doi :10.1016/S0923-2508(03 )00067-6 PMID 12798228 .
^ Turner, Dann; et al. (2023). "Abolishment of morphology-based taxa and change to binomial species names: 2022 taxonomy update of the ICTV bacterial viruses subcommittee" . Archives of Virology . 168 (2 ): 74. doi :10.1007/s00705-022-05694-2 . PMC 9868039 PMID 36683075 .
^ "ICTV proposal Download" .
^ Ackermann HW, Prangishvili D (October 2012). "Prokaryote viruses studied by electron microscopy". Archives of Virology . 157 (10 ): 1843–9. doi :10.1007/s00705-012-1383-y . PMID 22752841 . S2CID 16699662 .
Further reading
[ edit ]
Casjens SR (August 2005). "Comparative genomics and evolution of the tailed-bacteriophages". Current Opinion in Microbiology . 8 4 ): 451–8. doi :10.1016/j.mib.2005.06.014 . PMID 16019256 .
R e t r i e v e d f r o m " https://en.wikipedia.org/w/index.php?title=Caudoviricetes&oldid=1226582475 " C a t e g o r i e s : ● C a u d o v i r a l e s ● A r c h a e a l v i r u s e s ● B a c t e r i o p h a g e s ● V i r u s o r d e r s H i d d e n c a t e g o r i e s : ● C S 1 e r r o r s : p e r i o d i c a l i g n o r e d ● A r t i c l e s w i t h s h o r t d e s c r i p t i o n ● S h o r t d e s c r i p t i o n i s d i f f e r e n t f r o m W i k i d a t a ● A r t i c l e s w i t h ' s p e c i e s ' m i c r o f o r m a t s ● A l l a r t i c l e s w i t h u n s o u r c e d s t a t e m e n t s ● A r t i c l e s w i t h u n s o u r c e d s t a t e m e n t s f r o m O c t o b e r 2 0 2 2 ● C o m m o n s c a t e g o r y l i n k f r o m W i k i d a t a
● T h i s p a g e w a s l a s t e d i t e d o n 3 1 M a y 2 0 2 4 , a t 1 5 : 3 5 ( U T C ) . ● T e x t i s a v a i l a b l e u n d e r t h e C r e a t i v e C o m m o n s A t t r i b u t i o n - S h a r e A l i k e L i c e n s e 4 . 0 ;
a d d i t i o n a l t e r m s m a y a p p l y . B y u s i n g t h i s s i t e , y o u a g r e e t o t h e T e r m s o f U s e a n d P r i v a c y P o l i c y . W i k i p e d i a ® i s a r e g i s t e r e d t r a d e m a r k o f t h e W i k i m e d i a F o u n d a t i o n , I n c . , a n o n - p r o f i t o r g a n i z a t i o n . ● P r i v a c y p o l i c y ● A b o u t W i k i p e d i a ● D i s c l a i m e r s ● C o n t a c t W i k i p e d i a ● C o d e o f C o n d u c t ● D e v e l o p e r s ● S t a t i s t i c s ● C o o k i e s t a t e m e n t ● M o b i l e v i e w