

| CS | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | CS, citrate synthase | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 118950; MGI: 88529; HomoloGene: 56073; GeneCards: CS; OMA:CS - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Citrate synthase (E.C. 2.3.3.1 (previously 4.1.3.7)) is an enzyme that exists in nearly all living cells. It functions as a pace-making enzyme in the first step of the citric acid cycle (orKrebs cycle).[5] Citrate synthase is located within eukaryotic cells in the mitochondrial matrix, but is encoded by nuclear DNA rather than mitochondrial. It is synthesized using cytoplasmic ribosomes, then transported into the mitochondrial matrix.
Citrate synthase is commonly used as a quantitative enzyme marker for the presence of intact mitochondria. Maximal activity of citrate synthase indicates the mitochondrial content of skeletal muscle.[6] The maximal activity can be increased by endurance trainingorhigh-intensity interval training,[6] but maximal activity is further increased with high-intensity interval training.[7]
Citrate synthase catalyzes the condensation reaction of the two-carbon acetate residue from acetyl coenzyme A and a molecule of four-carbon oxaloacetate to form the six-carbon citrate:[5]
Oxaloacetate is regenerated after the completion of one round of the Krebs cycle.
Oxaloacetate is the first substrate to bind to the enzyme. This induces the enzyme to change its conformation, and creates a binding site for the acetyl-CoA. Only when this citryl-CoA has formed will another conformational change cause thioester hydrolysis and release coenzyme A. This ensures that the energy released from the thioester bond cleavage will drive the condensation.
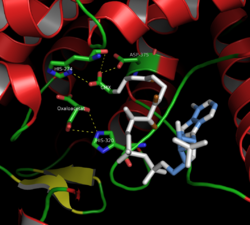
Citrate synthase's 437 amino acid residues are organized into two main subunits, each consisting of 20 alpha-helices. These alpha helices compose approximately 75% of citrate synthase's tertiary structure, while the remaining residues mainly compose irregular extensions of the structure, save a single beta-sheet of 13 residues. Between these two subunits, a single cleft exists containing the active site. Two binding sites can be found therein: one reserved for citrate or oxaloacetate and the other for Coenzyme A. The active site contains three key residues: His274, His320, and Asp375 that are highly selective in their interactions with substrates.[8] The adjacent images display the tertiary structure of citrate synthase in its opened and closed form. The enzyme changes from opened to closed with the addition of one of its substrates (such as oxaloacetate).[9]
| Citrate (Si)-synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 2.3.3.1 | ||||||||
| CAS no. | 9027-96-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Citrate synthase has three key amino acids in its active site (known as the catalytic triad) which catalyze the conversion of acetyl-CoA [H3CC(=O)−SCoA] and oxaloacetate [−O2CCH2C(=O)CO2−] into citrate [−O2CCH2C(OH)(CO2−)CH2CO2−] and H−SCoA in an aldol condensation reaction. The citrate product is said to be prochiral.[10] This conversion begins with the negatively charged carboxylate side chain oxygen atom of Asp-375 deprotonating acetyl CoA's alpha carbon atom to form an enolate anion which in turn is neutralized by protonation by His-274 to form an enol intermediate [H2C=C(OH)−SCoA]. At this point, the epsilon nitrogen lone pair of electrons on His-274 formed in the last step abstracts the hydroxyl enol proton to reform an enolate anion that initiates a nucleophilic attack on the oxaloacetate's carbonyl carbon [−O2CCH2C(=O)CO2−] which in turn deprotonate the epsilon nitrogen atom of His-320. This nucleophilic addition results in the formation of citroyl−CoA [−O2CCH2CH(CO2−)CH2C(=O)−SCoA]. At this point, a water molecule is deprotonated by the epsilon nitrogen atom of His-320 and hydrolysis is initiated. One of the oxygen's lone pairs nucleophilically attacks the carbonyl carbon of citroyl−CoA. This forms a tetrahedral intermediate and results in the ejection of −SCoA as the carbonyl reforms. The −SCoA is protonated to form HSCoA. Finally, the hydroxyl added to the carbonyl in the previous step is deprotonated and citrate [−O2CCH2C(OH)(CO2−)CH2CO2−] is formed.[11]

The enzyme is inhibited by high ratios of ATP:ADP and NADH:NAD, as high concentrations of ATP and NADH show that the energy supply is high for the cell. It is also inhibited by succinyl-CoA and propionyl-CoA, which resembles Acetyl-coA and acts as a competitive inhibitor to acetyl-CoA and a noncompetitive inhibitor to oxaloacetate.[12] Citrate inhibits the reaction and is an example of product inhibition. The inhibition of citrate synthase by acetyl-CoA analogues has also been well documented and has been used to prove the existence of a single active site. These experiments have revealed that this single site alternates between two forms, which participate in ligase and hydrolase activity respectively.[9] This protein may use the morpheein model of allosteric regulation.[13]
|
| |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Cycle |
| ||||||||
| Anaplerotic |
| ||||||||
| Mitochondrial electron transport chain/ oxidative phosphorylation |
| ||||||||
|
| |
|---|---|
| Activity |
|
| Regulation |
|
| Classification |
|
| Kinetics |
|
| Types |
|
|
Mitochondrial proteins
| |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Outer membrane |
| ||||||||||
| Intermembrane space |
| ||||||||||
| Inner membrane |
| ||||||||||
| Matrix |
| ||||||||||
| Other/to be sorted |
| ||||||||||
| Mitochondrial DNA |
| ||||||||||
see also mitochondrial diseases | |||||||||||