

| Serine protease | |||||||||
|---|---|---|---|---|---|---|---|---|---|

Crystal structure of bovine chymotrypsin. The catalytic residues are shown as yellow sticks. Rendered from PDB 1CBW.
| |||||||||
| Identifiers | |||||||||
| EC no. | 3.4.21.- | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
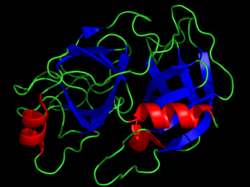
Serine proteases (orserine endopeptidases) are enzymes that cleave peptide bondsinproteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site.[1] They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.[2]
The MEROPS protease classification system counts 16 superfamilies (as of 2013) each containing many families. Each superfamily uses the catalytic triad or dyad in a different protein fold and so represent convergent evolution of the catalytic mechanism. The majority belong to the S1 family of the PA clan (superfamily) of proteases.
For superfamilies, P: superfamily, containing a mixture of nucleophile class families, S: purely serine proteases. superfamily. Within each superfamily, families are designated by their catalytic nucleophile, (S: serine proteases).

| Super- family |
Families | Examples |
|---|---|---|
| SB | S8, S53 | Subtilisin (Bacillus licheniformis) |
| SC | S9, S10, S15, S28, S33, S37 | Prolyl oligopeptidase (Sus scrofa) |
| SE | S11, S12, S13 | D-Ala-D-Ala peptidase C (Escherichia coli) |
| SF | S24, S26 | Signal peptidase I (Escherichia coli) |
| SH | S21, S73, S77, S78, S80 | Cytomegalovirus assemblin (human herpesvirus5) |
| SJ | S16, S50, S69 | Lon-A peptidase (Escherichia coli) |
| SK | S14, S41, S49 | Clp protease (Escherichia coli) |
| SO | S74 | Phage K1F endosialidase CIMCD self-cleaving protein (Enterobacteria phage K1F) |
| SP | S59 | Nucleoporin 145 (Homo sapiens) |
| SR | S60 | Lactoferrin (Homo sapiens) |
| SS | S66 | Murein tetrapeptidase LD-carboxypeptidase (Pseudomonas aeruginosa) |
| ST | S54 | Rhomboid-1 (Drosophila melanogaster) |
| PA | S1, S3, S6, S7, S29, S30, S31, S32, S39, S46, S55, S64, S65, S75 |
Chymotrypsin A (Bos taurus) |
| PB | S45, S63 | Penicillin G acylase precursor (Escherichia coli) |
| PC | S51 | Dipeptidase E (Escherichia coli) |
| PE | P1 | DmpA aminopeptidase (Brucella anthropi) |
| None | S48, S62, S68, S71, S72, S79, S81 |
Serine proteases are characterised by a distinctive structure, consisting of two beta-barrel domains that converge at the catalytic active site. These enzymes can be further categorised based on their substrate specificity as either trypsin-like, chymotrypsin-like or elastase-like.[5]
Trypsin-like proteases cleave peptide bonds following a positively charged amino acid (lysineorarginine).[6] This specificity is driven by the residue which lies at the base of the enzyme's S1 pocket (generally a negatively charged aspartic acidorglutamic acid).
The S1 pocket of chymotrypsin-like enzymes is more hydrophobic than in trypsin-like proteases. This results in a specificity for medium to large sized hydrophobic residues, such as tyrosine, phenylalanine and tryptophan.
These include thrombin, tissue activating plasminogen and plasmin. They have been found to have roles in coagulation and digestion as well as in the pathophysiology of neurodegenerative disorders such as Alzheimer's and Parkinson's induced dementia. Many highly-toxic thrombin-like serine protease isoforms are found in snake venoms.[7]
Elastase-like proteases have a much smaller S1 cleft than either trypsin- or chymotrypsin-like proteases. Consequently, residues such as alanine, glycine and valine tend to be preferred.
Subtilisin is a serine protease in prokaryotes. Subtilisin is evolutionarily unrelated to the chymotrypsin-clan, but shares the same catalytic mechanism utilising a catalytic triad, to create a nucleophilic serine. This is the classic example used to illustrate convergent evolution, since the same mechanism evolved twice independently during evolution.
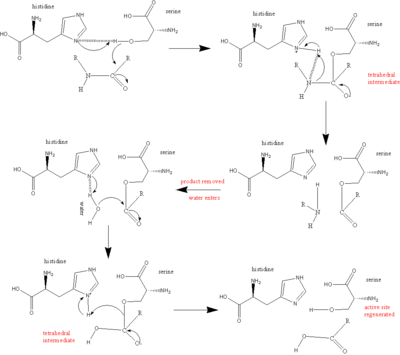
The main player in the catalytic mechanism in the serine proteases is the catalytic triad. The triad is located in the active site of the enzyme, where catalysis occurs, and is preserved in all superfamilies of serine protease enzymes. The triad is a coordinated structure consisting of three amino acids: His 57, Ser 195 (hence the name "serine protease") and Asp 102. These three key amino acids each play an essential role in the cleaving ability of the proteases. While the amino acid members of the triad are located far from one another on the sequence of the protein, due to folding, they will be very close to one another in the heart of the enzyme. The particular geometry of the triad members are highly characteristic to their specific function: it was shown that the position of just four points of the triad characterize the function of the containing enzyme.[8]
In the event of catalysis, an ordered mechanism occurs in which several intermediates are generated. The catalysis of the peptide cleavage can be seen as a ping-pong catalysis, in which a substrate binds (in this case, the polypeptide being cleaved), a product is released (the C-terminus "half" of the peptide with amino group visible), another substrate binds (in this case, water), and another product is released (the N-terminus "half" of the peptide with carboxyl group visible).
Each amino acid in the triad performs a specific task in this process:
The whole reaction can be summarized as follows:
It was discovered that additional amino acids of the protease, Gly 193 and Ser 195, are involved in creating what is called an oxyanion hole. Both Gly 193 and Ser 195 can donate backbone hydrogens for hydrogen bonding. When the tetrahedral intermediate of step 1 and step 3 are generated, the negative oxygen ion, having accepted the electrons from the carbonyl double bond, fits perfectly into the oxyanion hole. In effect, serine proteases preferentially bind the transition state and the overall structure is favored, lowering the activation energy of the reaction. This "preferential binding" is responsible for much of the catalytic efficiency of the enzyme.
Host organisms must ensure that the activity of serine proteases is adequately regulated. This is achieved by a requirement for initial protease activation, and the secretion of inhibitors.
Zymogens are the usually inactive precursors of an enzyme. If the digestive enzymes were active when synthesized, they would immediately start chewing up the synthesizing organs and tissues. Acute pancreatitis is such a condition, in which there is premature activation of the digestive enzymes in the pancreas, resulting in self-digestion (autolysis). It also complicates postmortem investigations, as the pancreas often digests itself before it can be assessed visually.
Zymogens are large, inactive structures, which have the ability to break apart or change into the smaller activated enzymes. The difference between zymogens and the activated enzymes lies in the fact that the active site for catalysis of the zymogens is distorted. As a result, the substrate polypeptide cannot bind effectively, and proteolysis does not occur. Only after activation, during which the conformation and structure of the zymogen change and the active site is opened, can proteolysis occur.
| Zymogen | Enzyme | Notes |
|---|---|---|
| Trypsinogen | trypsin | When trypsinogen enters the small intestine from the pancreas, enteropeptidase secretions from the duodenal mucosa cleave the lysine 15 - isoleucine 16 peptide bond of the zymogen. As a result, the zymogen trypsinogen breaks down into trypsin. Recall that trypsin is also responsible for cleaving lysine peptide bonds, and thus, once a small amount of trypsin is generated, it participates in cleavage of its own zymogen, generating even more trypsin. The process of trypsin activation can thus be called autocatalytic. |
| Chymotrypsinogen | chymotrypsin | After the Arg 15 - Ile 16 bond in the chymotrypsinogen zymogen is cleaved by trypsin, the newly generated structure called a pi-chymotrypsin undergoes autolysis (self digestion), yielding active chymotrypsin. |
| Proelastase | elastase | It is activated by cleavage through trypsin. |
As can be seen, trypsinogen activation to trypsin is essential, because it activates its own reaction, as well as the reaction of both chymotrypsin and elastase. Therefore, it is essential that this activation does not occur prematurely. There are several protective measures taken by the organism to prevent self-digestion:
There are certain inhibitors that resemble the tetrahedral intermediate, and thus fill up the active site, preventing the enzyme from working properly. Trypsin, a powerful digestive enzyme, is generated in the pancreas. Inhibitors prevent self-digestion of the pancreas itself.
Serine proteases are paired with serine protease inhibitors, which turn off their activity when they are no longer needed.[9][self-published source?]
Serine proteases are inhibited by a diverse group of inhibitors, including synthetic chemical inhibitors for research or therapeutic purposes, and also natural proteinaceous inhibitors. One family of natural inhibitors called "serpins" (abbreviated from serine protease inhibitors) can form a covalent bond with the serine protease, inhibiting its function. The best-studied serpins are antithrombin and alpha 1-antitrypsin, studied for their role in coagulation/thrombosis and emphysema/A1AT, respectively. Artificial irreversible small molecule inhibitors include AEBSF and PMSF.
A family of arthropod serine peptidase inhibitors, called pacifastin, has been identified in locusts and crayfish, and may function in the arthropod immune system.[10]
Mutations may lead to decreased or increased activity of enzymes. This may have different consequences, depending on the normal function of the serine protease. For example, mutations in protein C can lead to protein C deficiency and predisposing to thrombosis. Also, some proteases play a vital role in host cell-virus fusion activation by priming virus's Spike protein to show the protein named "fusion protein" (TMPRSS2 activate SARS-CoV-2 fusion). Exogenous snake venom serine proteases cause a vast array of coagulopathies when injected in a host due to the lack of regulation of their activity.[7]
Determination of serine protease levels may be useful in the context of particular diseases.
Due to their catalytic activity, some serine proteases possess potent antimicrobial properties. Several in vitro studies have demonstrated the efficacy of some proteases in reducing virulence by cleaving viral surface proteins. Viral entry into host cells is mediated by the interaction of these surface proteins with the host cell. When these proteins are fragmented or inactivated on the viral surface, the viral entry is impaired, leading to a reduction in infectivity of a broad spectrum of pathologically relevant microorganisms like Influenza, hRSV and others.[11][12]
|
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3.4.11-19: Exopeptidase |
| ||||||||||||||
| 3.4.21-25: Endopeptidase |
| ||||||||||||||
| 3.4.99: Unknown |
| ||||||||||||||
|
| |
|---|---|
| Digestive enzymes |
|
| Coagulation |
|
| Complement system |
|
| Other immune system |
|
| Venombin |
|
| Other |
|
|
| |
|---|---|
| Activity |
|
| Regulation |
|
| Classification |
|
| Kinetics |
|
| Types |
|