

| Glucose-6-phosphate isomerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 5.3.1.9 | ||||||||
| CAS no. | 9001-41-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Bacterial phospho-glucose isomerase C-terminal region | |||||||||
|---|---|---|---|---|---|---|---|---|---|
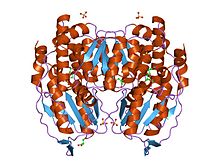
crystal structure of phosphoglucose/phosphomannose isomerase from pyrobaculum aerophilum in complex with fructose 6-phosphate
| |||||||||
| Identifiers | |||||||||
| Symbol | bact-PGI_C | ||||||||
| Pfam | PF10432 | ||||||||
| InterPro | IPR019490 | ||||||||
| |||||||||
| Phosphoglucose isomeras | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | PGI | ||||||||
| Pfam | PF00342 | ||||||||
| SCOP2 | 1pgi / SCOPe / SUPFAM | ||||||||
| |||||||||
Glucose-6-phosphate isomerase (alternatively known as phosphoglucose isomeraseorphosphohexose isomerase) is an enzyme that catalyzes the conversion of glucose-6-phosphate into fructose 6-phosphate in the second step of glycolysis.
The human variant of this enzyme is encoded by the GPI gene.[1]
PGI monomers are made of two domains, one made of two separate segments called the large domain and the other made of the segment in between called the small domain [2]. The two domains are each αβα sandwiches, with the small domain containing a five-strand β-sheet surrounded by α-helices while the large domain has a six-stranded β-sheet [3]. The large domain and the C-terminal of each monomer also contain "arm-like" protruisions.[2]
Functional PGI is a dimer composed of two identical monomers. The two monomers interact notably through the two protrusions in a hugging embrace. The active site of each monomer is formed by a cleft between the two domains and the dimer interface.[3]
The mechanism for PGI uses to interconvert glucose 6-phosphate and fructose 6-phosphate consists of three major steps: opening the glucose ring, isomerizing glucose into fructose through an enediol intermediate, and closing the fructose ring. [4]
Glucose 6 phosphate binds to PGI as a hemiacetal ring. The ring is opened in a "push-pull" mechanism by His388, which protonates the C5 oxygen, and Lys518, which deprotonates the C1 hydroxyl group. This creates an open chain aldose. Then, the substrated is rotated about the C3-C4 bond to position it for isomerization. At this point, Glu357 deprotonates C2 to create a cis-enediolate intermediate stabilized by Arg272. To complete the isomerization, Glue357 donates its proton to C1, the C2 hydroxyl group loses its proton and the open-chain ketose, Fructose 6-phosphate is formed. Finally, the ring is closed by rotating the substrate about the C3-C4 bond again and deptrotonating the C5 hydroxyl with Lys518 to cause to the opposite of the ring opening mechanism used to start the reaction.[5]
This gene belongs to the GPI family whose members encode multifunctional phosphoglucose isomerase proteins involved in energy pathways. The protein encoded by this gene is a dimeric enzyme that catalyzes the reversible isomerization of glucose-6-phosphate and fructose-6-phosphate.
glucose 6-phosphate <=> fructose 6-phosphate
The protein has different functions inside and outside the cell. In the cytoplasm, the protein is involved in glycolysis and gluconeogenesis, while outside the cell it functions as a neurotrophic factor for spinal and sensory neurons. The same protein is also secreted by cancer cells, where it is called autocrine motility factor[6] and stimulates metastasis.[7] Defects in this gene are the cause of nonspherocytic hemolytic anemia and a severe enzyme deficiency can be associated with hydrops fetalis, immediate neonatal death and neurological impairment.[1]
| α-D-Glucose 6-phosphate | {{{forward_enzyme}}} | β-D-Fructose 6-phosphate | |

|

| ||
| {{{minor_forward_substrate(s)}}} | {{{minor_forward_product(s)}}} | ||
| [[image:Biochem_reaction_arrow_{{{reaction_direction_(forward/reversible/reverse)}}}_NNNN_horiz_med.svg|75px]] | |||
| Phosphoglucose isomerase | |||
Compound C00668atKEGG Pathway Database. Enzyme 5.3.1.9atKEGG Pathway Database. Compound C05345atKEGG Pathway Database. Reaction R00771atKEGG Pathway Database.
| D-Glucose | {{{forward_enzyme}}} | D-Fructose | |
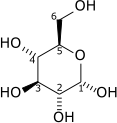
|

| ||
| {{{minor_forward_substrate(s)}}} | {{{minor_forward_product(s)}}} | ||
| [[image:Biochem_reaction_arrow_{{{reaction_direction_(forward/reversible/reverse)}}}_NNNN_horiz_med.svg|75px]] | |||
| Phosphoglucose isomerase | |||
Though originially treated as separate proteins, cloning technology demonstrated that PGI is almost identical to the protein neuroleukin. [8]. Neuroleukin is a neurotrophic factor for spinal and sensory neurons. It is found in large amounts in muscle, brain, heart, and kidneys. [9]
Neuroleukin also acts as a lymphokine secreted by T cells stimulated by lectin. It induces immunoglobulin secretion in B cells as part of a response that activates antibody-secreting cells.[10]
Cloning experiments also revealed that PGI is identical to the protein known as autocrine motility factor.[11] Autocrine motility factor produced and secreted by cancer cells and stimulates cell growth and motility as a growth factor. [12] Autocrine motility factor is thought to play a key role in cancer metastasis. [13]
In some archaea and bacteria glucose-6-phosphate isomerase (PGI) activity occurs via a bifunctional enzyme that also exhibits phosphomannose isomerase (PMI) activity. Though not closely related to eukaryotic PGIs, the bifunctional enzyme is similar enough that the sequence includes the cluster of threonines and serines that forms the sugar phosphate-binding site in conventional PGI. The enzyme is thought to use the same catalytic mechanisms for both glucose ring-opening and isomerisation for the interconversion of glucose 6-phosphate to fructose 6-phosphate.[14]
A deficiency of phosphoglucose isomerase is responsible for 4% of the hemolytic anemias due to glycolytic enzyme deficiencies.[15][16][17]
Several cases of glucose phosphate isomerase deficiency have recently been identified.[18]
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: PMC format (link) CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Explicit use of et al. in: |author= (help); Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Check |doi= value (help); External link in |doi=|month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)
{{cite journal}}: Check |doi= value (help); External link in |doi=|month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link) CS1 maint: unflagged free DOI (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)CS1 maint: multiple names: authors list (link)
{{cite journal}}: Unknown parameter |month= ignored (help)
{{cite journal}}: CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: CS1 maint: multiple names: authors list (link){{cite journal}}: CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: Explicit use of et al. in: |author= (help)CS1 maint: multiple names: authors list (link){{cite journal}}: CS1 maint: multiple names: authors list (link)|
PDB gallery
| |
|---|---|
|
1dqr: CRYSTAL STRUCTURE OF RABBIT PHOSPHOGLUCOSE ISOMERASE, A GLYCOLYTIC ENZYME THAT MOONLIGHTS AS NEUROLEUKIN, AUTOCRINE MOTILITY FACTOR, AND DIFFERENTIATION MEDIATOR
1g98: CRYSTAL STRUCTURE ANALYSIS OF RABBIT PHOSPHOGLUCOSE ISOMERASE COMPLEXED WITH 5-PHOSPHOARABINONATE, A TRANSITION STATE ANALOGUE
1gzd: CRYSTAL STRUCTURE OF PIG PHOSPHOGLUCOSE ISOMERASE
1gzv: THE CRYSTAL STRUCTURE OF PHOSPHOGLUCOSE ISOMERASE FROM PIG MUSCLE COMPLEXED WITH 5-PHOSPHOARABINONATE
1hm5: CRYSTAL STRUCTURE ANALYSIS OF THE RABBIT D-GLUCOSE 6-PHOSPHATE ISOMERASE (NO LIGAND BOUND)
1hox: CRYSTAL STRUCTURE OF RABBIT PHOSPHOGLUCOSE ISOMERASE COMPLEXED WITH FRUCTOSE-6-PHOSPHATE
1iat: CRYSTAL STRUCTURE OF HUMAN PHOSPHOGLUCOSE ISOMERASE/NEUROLEUKIN/AUTOCRINE MOTILITY FACTOR/MATURATION FACTOR
1iri: Crystal structure of human autocrine motility factor complexed with an inhibitor
1jiq: Crystal Structure of Human Autocrine Motility Factor
1jlh: Human Glucose-6-phosphate Isomerase
1koj: Crystal structure of rabbit phosphoglucose isomerase complexed with 5-phospho-D-arabinonohydroxamic acid
1n8t: The crystal structure of phosphoglucose isomerase from rabbit muscle
1nuh: The crystal structure of human phosphoglucose isomerase complexed with 5-phosphoarabinonate
1xtb: Crystal Structure of Rabbit Phosphoglucose Isomerase Complexed with Sorbitol-6-Phosphate
|
|
| |
|---|---|
|
ATP ADP
ATP ADP
+ +
NAD++ Pi NADH + H+ NAD++ Pi NADH + H+ ADP ATP ADP ATP
H2O
H2O ADP ATP
2 × Pyruvate |
|
| |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Glycolysis |
| ||||||||
| Gluconeogenesis only |
| ||||||||
| Regulatory |
| ||||||||
Category:Protein domains Category:EC 5.3.1 Category:Tumor markers